Contents
- 1 FAQ Section 6: Postprocessing
- 1.1 Postprocessing of Lagrangian particles
- 1.2 Postprocessing of simulations on multiple mesh-regions
- 1.3 Post-processing multiphase results
- 1.4 Artificial particle tracking in ParaView
- 1.5 Visual mesh diagnosis in ParaView
- 1.6 Using the internal .foam reader in ParaView
- 1.7 Post-processing sub-domains (parallel/decomposed cases) in ParaView
- 1.8 I'm using Ubuntu, where is the main menu in ParaView?
- 1.9 Using the internal reader in ParaView cannot load unknown boundary conditions?
1 FAQ Section 6: Postprocessing
Working with the results
1.1 Postprocessing of Lagrangian particles
The following subsections describe the basics on how to post-process by Using foamToVTK and by Using paraFoam.
Beyond this, have a look into:
- the tutorial page HowTo Post-process results for particle based methods.
- the tips page Tip Paraview for Lagrangian Data
1.1.1 Using foamToVTK
With the latest OpenFOAM and ParaView versions, it's possible to post-process Lagrangian data directly in ParaView. However, if for some reason you cannot open the OpenFOAM case directly in ParaView, the workaround is to use foamToVTK and ParaView's existing ability to handle point data.
Follow these steps:
- Run the foamToVTK post-processing utility. As with all OpenFOAM programs, it requires at a minimum, the root and case paths.
- Read in your Eulerian data, if you wish.
- Read the Lagrangian data separately and 'Glyph' it. Glyphs are how ParaView represents point data. Usually, the sphere glyph is the most appropriate.
The glyphs used to represent particles can be colored and sized to reflect the data associated with the particle. If you cannot see your particles, check that indeed you have represented them as glyphs and that the size scaling you have chosen is adequately large.
1.1.2 Using paraFoam
First you execute paraFoam as usual but you select only in the data to be load the fields and the mesh of the continuous phase.
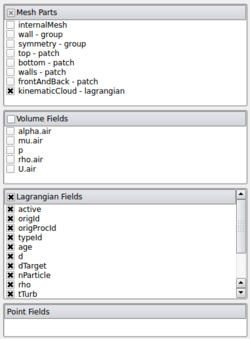
Then you click on the Open button (or File -> Open in the menu). You select the case.OpenFOAM file created by paraFoam. So you can load again the data of the case. But this time, you select:
|
Then click on the Apply button.
To visualize the particle, use the Glyph filter (cf. end of previous paragraph).
Remark: Should work as of OpenFOAM-1.6 and ParaView 3.6.1.
1.2 Postprocessing of simulations on multiple mesh-regions
Two possibilites to do this are described on How To Postprocess Multiple Regions
1.3 Post-processing multiphase results
OpenFOAM handles multiphase simulations with multiphase flow solver through using a single field that represents the phase present at a certain cell, face or vertex. The field is usually named alpha or alpha1. For more information on how to analyse this field, see this page: How to post-process multiphase results
1.4 Artificial particle tracking in ParaView
This question refers to non-Lagrangian simulations, such as the ones done with icoFoam and simpleFoam, where the objective is to calculate the path that a particle would have if placed inside the flow domain. This is explained in some detail in the following forum post: do particle tracking in paraview? - post #3
1.5 Visual mesh diagnosis in ParaView
People usually tend to use the filters Slice and Clip to diagnose the interior mesh of a case. Although it's an idea in the right direction (wanting to look inside the mesh), it will not give you the best results for assessing if the mesh is OK or not. This is because these filters require that the polyhedral cells in the mesh be decomposed to tetrahedral cells and the resulting surfaces will all be composed by triangles.
The correct way to diagnose the cell structure mesh in ParaView is to use the filter Extract Cells by Region. In addition, you should load the mesh for the case without polyhedral decomposition:
| To the right is an example of selecting the Use VTKPolyhedron option in the official reader, which usually appears when the file extension .OpenFOAM is used. Notice mesh representation shown in 3D by the filter Extract Cells by Region.
The file extension .OpenFOAM is usually used automatically when you simply run: paraFoam |
|
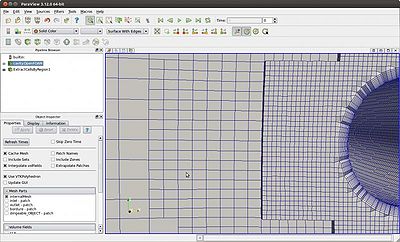
| To the right is another example of how to not select the Decompose polyhedra option when the file extension .foam is used. Notice mesh representation shown in 3D by the filter Extract Cells by Region.
The file extension .foam is usually used automatically when you run:
|
|
If for some reason you cannot use the official or internal readers in ParaView, you can use foamToVTK to export the mesh without decomposing cells, by using the -poly option:
foamToVTK -poly |
(no image available) |
In addition to visual inspection, ParaView provides a Mesh Quality filter that is useful for identifying degenerate elements. It assigns a floating-point "quality" to each cell (many of these quality metrics are related to the condition number of the Jacobian which is used during numerical integration). Once the quality has been computed for each cell, you can use the Threshold filter to isolate only those cells that misbehave. Rendering the original mesh with transparency turned on (or in wireframe mode) will show the thresholded cells even if they are inside the mesh and would not otherwise be visible.
1.6 Using the internal .foam reader in ParaView
Depending on the OpenFOAM version or fork you're using, the paraFoam script is able to use the internal .foam reader:
Otherwise, at least in most OpenFOAM versions, the default file extension used is .OpenFOAM, which usually is associated to the official OpenFOAM plug-in for ParaView, which needs to be built from OpenFOAM's source code.
1.7 Post-processing sub-domains (parallel/decomposed cases) in ParaView
Note: The following description assumes you've already studied the topic Using the internal .foam reader in ParaView.
There are a few ways to post-process cases that have been decomposed into sub-domains and that have been run in parallel, which as listed as follows:
- Reconstructing the case
- Sometimes the only accurate way of post-processing a case is to reconstruct it first (the opposite of decomposing). There are 2 main types of reconstruction:
- Using reconstructParMesh is only needed when a new mesh was generated in parallel. Usually this happens when using snappyHexMesh in parallel.
- Using reconstructPar should work on all other situations, where it will reconstruct the fields and dynamic meshes.
- Using foamToVTK in parallel
- When all fails, i.e. if you're not able to open the case directly in ParaView for whatever reason, you can still run foamToVTK in parallel, the exact same way you executed the solver or mesher in parallel. For example, if you ran the solver like this:
-
foamJob -p simpleFoam
- Then you can also run foamToVTK like this:
-
foamJob -p foamToVTK
- Then you can open the respective files for the respective sub-domains that are related to each processor* folder.
- Opening one processor* folder at a time
- paraFoam can be used for opening directly a sub-domain, by running something like this:
-
paraFoam -case processor0
- Using the internal reader
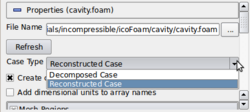
- As mentioned before, the topic Using the internal .foam reader in ParaView already addresses the fact that the internal reader can be used in many cases. What might not have been mentioned, is that there is an option to open the decomposed case, instead of using the default option of opening the reconstructed case, as shown in the image on the right.
- The idea here is that this will allow you to post-process the case without having to first reconstruct the case.
That's a feature of Ubuntu's Unity, where the menu is on the bar on the top of the screen. To see the menu, follow these steps:
- Look at the top bar on your screen.
- Move your mouse cursor over that bar...
- Aim a bit to the middle/middle-left of the screen...
- See the menu appearing there?
1.9 Using the internal reader in ParaView cannot load unknown boundary conditions?
There are several special boundary conditions in OpenFOAM that do not write/use the entry value in a boundary condition, which leads to the internal reader that opens with the file extension .foam to fail to load the fields.
- Workarounds
- Artificially add the value entry with Template:ChangeDictionary.
- Use foamToVTK to export the results.
- Use the official OpenFOAM reader that is built with OpenFOAM.
| Faqdescription | Working with the results + |
| Faqname | Postprocessing + |
| Faqnumber | 6 + |