OverPimpleDyMFoam
OverPimpleDyMFoam
Transient solver for incompressible flow of Newtonian fluids on a moving mesh using the PIMPLE (merged PISO-SIMPLE) algorithm. Turbulence modelling is generic, i.e. laminar, RAS or LES may be selected.
Contents
1 Solution Strategy
The solver follows a segregated solution strategy. This means that the equations for each variable characterizing the system (the velocity 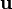 , the pressure
, the pressure 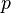 and the variables characterizing turbulence) is solved sequentially
and the solution of the preceding equations is inserted in the subsequent equation. The non-linearity appearing in the momentum equation (the face flux
and the variables characterizing turbulence) is solved sequentially
and the solution of the preceding equations is inserted in the subsequent equation. The non-linearity appearing in the momentum equation (the face flux
 which is a function of the velocity) is resolved by computing it from the velocity and pressure values of the preceding iteration. The dependence from
the pressure is introduced to avoid a decoupling between the momentum and pressure equations and hence the appearance of high frequency oscillation in the solution (check board effect).
The first equation to be solve is the momentum equation. It delivers a velocity field
which is a function of the velocity) is resolved by computing it from the velocity and pressure values of the preceding iteration. The dependence from
the pressure is introduced to avoid a decoupling between the momentum and pressure equations and hence the appearance of high frequency oscillation in the solution (check board effect).
The first equation to be solve is the momentum equation. It delivers a velocity field  which is in general not divergence free, i.e.
it does not satisfy the continuity equation. After that the momentum and the continuity equations are used to construct an equation for the pressure. The
aim is to obtain a pressure field
which is in general not divergence free, i.e.
it does not satisfy the continuity equation. After that the momentum and the continuity equations are used to construct an equation for the pressure. The
aim is to obtain a pressure field  , which, if inserted in the momentum equation, delivers a divergence free velocity field
, which, if inserted in the momentum equation, delivers a divergence free velocity field 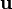 .
After correcting the velocity field, the equations for turbulence are solved.
The above iterative solution procedure is repeated until convergence.
.
After correcting the velocity field, the equations for turbulence are solved.
The above iterative solution procedure is repeated until convergence.
The overset method allows to solve the governing equaiton on a set of disjoint meshes, i.e., the meshes are not connected over faces. The coupling between the differnt meshes is done over an implicit interpolation.
The source code can be found in overPimpleDyMFoam.C
/*---------------------------------------------------------------------------*\ ========= | \\ / F ield | OpenFOAM: The Open Source CFD Toolbox \\ / O peration | \\ / A nd | www.openfoam.com \\/ M anipulation | ------------------------------------------------------------------------------- Copyright (C) 2011-2016 OpenFOAM Foundation Copyright (C) 2016-2018 OpenCFD Ltd. ------------------------------------------------------------------------------- License This file is part of OpenFOAM. OpenFOAM is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version. OpenFOAM is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details. You should have received a copy of the GNU General Public License along with OpenFOAM. If not, see <http://www.gnu.org/licenses/>. Application overPimpleDyMFoam Group grpIncompressibleSolvers grpMovingMeshSolvers Description Transient solver for incompressible flow of Newtonian fluids on a moving mesh using the PIMPLE (merged PISO-SIMPLE) algorithm. Turbulence modelling is generic, i.e. laminar, RAS or LES may be selected. \*---------------------------------------------------------------------------*/ #include "fvCFD.H" #include "dynamicFvMesh.H" #include "singlePhaseTransportModel.H" #include "turbulentTransportModel.H" #include "pimpleControl.H" #include "fvOptions.H" #include "cellCellStencilObject.H" #include "zeroGradientFvPatchFields.H" #include "localMin.H" #include "interpolationCellPoint.H" #include "transform.H" #include "fvMeshSubset.H" #include "oversetAdjustPhi.H" // * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * // int main(int argc, char *argv[]) { argList::addNote ( "Transient solver for incompressible, turbulent flow" " on a moving mesh." ); #include "postProcess.H" #include "setRootCaseLists.H" #include "createTime.H" #include "createDynamicFvMesh.H" #include "initContinuityErrs.H" pimpleControl pimple(mesh); #include "createFields.H" #include "createUf.H" #include "createMRF.H" #include "createFvOptions.H" #include "createControls.H" #include "CourantNo.H" #include "setInitialDeltaT.H" turbulence->validate(); // * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * * // Info<< "\nStarting time loop\n" << endl; while (runTime.run()) { #include "readControls.H" #include "CourantNo.H" #include "setDeltaT.H" ++runTime; Info<< "Time = " << runTime.timeName() << nl << endl; bool changed = mesh.update(); if (changed) { #include "setCellMask.H" #include "setInterpolatedCells.H" surfaceScalarField faceMaskOld ( localMin<scalar>(mesh).interpolate(cellMask.oldTime()) ); // Zero Uf on old faceMask (H-I) Uf *= faceMaskOld; // Update Uf and phi on new C-I faces Uf += (1-faceMaskOld)*fvc::interpolate(U); phi = mesh.Sf() & Uf; // Zero phi on current H-I surfaceScalarField faceMask ( localMin<scalar>(mesh).interpolate(cellMask) ); phi *= faceMask; } if (mesh.changing() && correctPhi) { // Calculate absolute flux from the mapped surface velocity #include "correctPhi.H" } // Make the flux relative to the mesh motion fvc::makeRelative(phi, U); if (mesh.changing() && checkMeshCourantNo) { #include "meshCourantNo.H" } // --- Pressure-velocity PIMPLE corrector loop while (pimple.loop()) { #include "UEqn.H" // --- Pressure corrector loop while (pimple.correct()) { #include "pEqn.H" } if (pimple.turbCorr()) { laminarTransport.correct(); turbulence->correct(); } } runTime.write(); runTime.printExecutionTime(Info); } Info<< "End\n" << endl; return 0; } // ************************************************************************* //
2 Equations
The equation of motion used for moving meshes are written in the Arbitrary-Euler-Lagrange (ALE) formulation. This formulation is one of the most popular if morphing meshes are used to describe the solid body deformation or displacement [1]. If the overset method is used, this formulation avoids to formulate the equation of motion in multiple frames of references [2]. For the derivation of the equation see [3] [4]. The continuity and momentum equation read in this form:
-
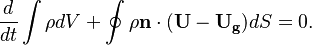 | (1) |
-
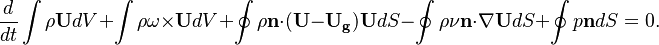 | (1) |
In the above equations $\U$ is the fluid velocity, 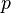 the pressure,
the pressure, 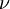 the molecular viscosity and
the molecular viscosity and 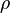 the density. The only difference to the equation of motion in a fixed mesh is the introduction of a new variable, i.e. the grid velocity
the density. The only difference to the equation of motion in a fixed mesh is the introduction of a new variable, i.e. the grid velocity 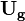 . In order to determine the grid velocity the space conservation law (SCL) has to be solved (see also [5] [6])
. In order to determine the grid velocity the space conservation law (SCL) has to be solved (see also [5] [6])
-
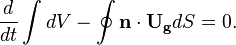 | (1) |
In order to not introduce additional mass conservation errors, the temporal discretisation of the SCL should be the same as used in the other conservation equation (see also [7] [8])
OpenFOAM uses a slightly different continuity equation (which is used to derive the equation for the pressure). By inserting the mass conservation law into the continuity equation we get for the spatial case of incompressible case:
-
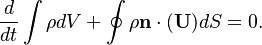 | (1) |
It is the same as the one for fixed grids. Note this transformation is valid only for incompressible flows.
2.1 Momentum Equation
In this section we will analyze the momentum equation implemented in the solver a bit more in detail. Es already seen above, the momentum equation reads:
-
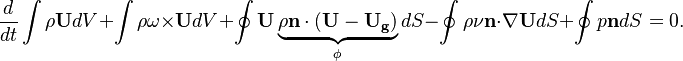 | (1) |
 is the mass flux over the surface
is the mass flux over the surface 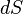 .
. 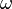 is the angular velocity of the rotating frame of reference. We see that the momentum equation is essentially the same as the one in fixed control volume. See, e.g. https://openfoamwiki.net/index.php/SimpleFoam. The only difference is how the phase flux
is the angular velocity of the rotating frame of reference. We see that the momentum equation is essentially the same as the one in fixed control volume. See, e.g. https://openfoamwiki.net/index.php/SimpleFoam. The only difference is how the phase flux  is calculated. In a fixed mesh, only the velocity in the inertial coordinate system
is calculated. In a fixed mesh, only the velocity in the inertial coordinate system 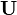 is used to calculate it, while for the overset mesh the difference between the velocity in the inertial coordinate system <math< \mathbf{U} </math> and the grid velocity
is used to calculate it, while for the overset mesh the difference between the velocity in the inertial coordinate system <math< \mathbf{U} </math> and the grid velocity 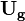 is used.
is used.
The source code of the momentum equation is shown above. We see that in front of the pressure equation the field cellMask is applied. It is 0 for hole cells and 1 elsewhere. This means that for background cells which are covered by a solid structure the pressure gradient term does not give any contribution to the momentum equation.
// Solve the Momentum equation MRF.correctBoundaryVelocity(U); tmp<fvVectorMatrix> tUEqn ( fvm::ddt(U) + fvm::div(phi, U) + MRF.DDt(U) + turbulence->divDevReff(U) == fvOptions(U) ); fvVectorMatrix& UEqn = tUEqn.ref(); UEqn.relax(); fvOptions.constrain(UEqn); if (pimple.momentumPredictor()) { solve(UEqn == -cellMask*fvc::grad(p)); fvOptions.correct(U); }
2.2 Pressure Equation
Since the continuity equation for incompressible flow written in an AEL (arbitrary Euler-Lagrange) formulation is the same as for fixed grid (see previous section), also the pressure equation is basically the same. For this reason the derivation of the pressure equation is omitted. For a detailed derivation see https://openfoamwiki.net/index.php/SimpleFoam and the references therein. In this section we will concentrate on the peculiarities of the pressure equation introduces for the overset method.
The code looks like:
// Option 1: interpolate rAU, do not block out rAU on blocked cells volScalarField rAU("rAU", 1.0/UEqn.A()); mesh.interpolate(rAU); // Option 2: do not interpolate rAU but block out rAU //surfaceScalarField rAUf("rAUf", fvc::interpolate(blockedCells*rAU)); // Option 3: do not interpolate rAU but zero out rAUf on faces on holes // But what about: // // H // H I C C C C // H // surfaceScalarField rAUf("rAUf", fvc::interpolate(rAU)); volVectorField H("H", UEqn.H()); volVectorField HbyA("HbyA", U); HbyA = constrainHbyA(rAU*H, U, p); if (massFluxInterpolation) { #include "interpolatedFaces.H" } if (runTime.outputTime()) { H.write(); rAU.write(); HbyA.write(); } if (pimple.nCorrPISO() <= 1) { tUEqn.clear(); } phiHbyA = fvc::flux(HbyA); if (ddtCorr) { surfaceScalarField faceMaskOld ( localMin<scalar>(mesh).interpolate(cellMask.oldTime()) ); phiHbyA += rAUf*faceMaskOld*fvc::ddtCorr(U, Uf); } MRF.makeRelative(phiHbyA); // WIP if (p.needReference()) { fvc::makeRelative(phiHbyA, U); adjustPhi(phiHbyA, U, p); fvc::makeAbsolute(phiHbyA, U); } if (adjustFringe) { fvc::makeRelative(phiHbyA, U); oversetAdjustPhi(phiHbyA, U); fvc::makeAbsolute(phiHbyA, U); } while (pimple.correctNonOrthogonal()) { fvScalarMatrix pEqn ( fvm::laplacian(rAUf, p) == fvc::div(phiHbyA) ); pEqn.setReference(pRefCell, pRefValue); pEqn.solve(mesh.solver(p.select(pimple.finalInnerIter()))); if (pimple.finalNonOrthogonalIter()) { phi = phiHbyA - pEqn.flux(); // option 2: // rAUf*fvc::snGrad(p)*mesh.magSf(); } } #include "continuityErrs.H" // Explicitly relax pressure for momentum corrector p.relax(); volVectorField gradP(fvc::grad(p)); // Option 2: zero out velocity on blocked out cells //U = HbyA - rAU*cellMask*gradP; // Option 3: zero out velocity on blocked out cells // This is needed for the scalar Eq (k,epsilon, etc) // which can use U as source term U = cellMask*(HbyA - rAU*gradP); U.correctBoundaryConditions(); fvOptions.correct(U); { Uf = fvc::interpolate(U); surfaceVectorField n(mesh.Sf()/mesh.magSf()); Uf += n*(phi/mesh.magSf() - (n & Uf)); } // Make the fluxes relative to the mesh motion fvc::makeRelative(phi, U); surfaceScalarField faceMask ( localMin<scalar>(mesh).interpolate(cellMask) ); phi *= faceMask;
Looking at the source code we see that we have two code passages which try to correct the mass flux imbalance produced by the overset interpolation:
if (massFluxInterpolation) { #include "interpolatedFaces.H" }
and
if (adjustFringe) { fvc::makeRelative(phiHbyA, U); oversetAdjustPhi(phiHbyA, U); fvc::makeAbsolute(phiHbyA, U); }
They are activated by setting the switches adjustFringe and massFluxInterpolation in the PIMPLE subdict of the fvSolution dictionary to yes.
2.2.1 AdjustFringe
If the adjustFringe switch is set to yes, the code loops over all faces. After that it checks if the faces are at the border of a calculated and an interpolated cell. If yes, the mass flux pointing into the calculated cell and the mass flux pointing outside the cell is summed over all faces in a mesh zone. Then a correction factor is calculated which is the ratio between sum over all mesh zone faces of the positive mass flux going over the boarder of calculated and interpolated cells and the negative mass flux. This correction factor is applied to all faces which are at the boarder of calculated and interpolated cells. In this way the global mass balance over a mesh zone is enforced. Note that a mesh zone is a topological connected region of the overset mesh.
The first part of the code is shown below. The flux is defined pointing from a calculated cell towards an interpolated cells. The code fist loops over all internal faces and calculates the mass flux going into the calculated cell bordering a interpolated cell (negative) and outside the calculated cell bordering a interpolated (positive). The same is done by looping over the boundary faces. In order to determine the type of cells on the neighbor processor the functionsyncTools::swapBoundaryCellList(mesh, cellTypes, neiCellTypes);
is called.
forAll(own, facei)
{
label zonei = zoneID[own[facei]]; // note:own and nei in same zone
label ownType = cellTypes[own[facei]];
label neiType = cellTypes[nei[facei]];
bool ownCalc =
(ownType == cellCellStencil::CALCULATED)
&& (neiType == cellCellStencil::INTERPOLATED);
bool neiCalc =
(ownType == cellCellStencil::INTERPOLATED)
&& (neiType == cellCellStencil::CALCULATED);
if (ownCalc || neiCalc)
{
// Calculate flux w.r.t. calculated cell
scalar flux = phi[facei];
if (neiCalc)
{
flux = -flux;
}
if (flux < 0.0)
{
massIn[zonei] -= flux;
}
else
{
adjustableMassOut[zonei] += flux;
}
}
}
}
// Check all coupled faces on the outside of interpolated cells
labelList neiCellTypes;
syncTools::swapBoundaryCellList(mesh, cellTypes, neiCellTypes);
{
forAll(bphi, patchi)
{
const fvPatchVectorField& Up = U.boundaryField()[patchi];
const fvsPatchScalarField& phip = bphi[patchi];
const labelUList& fc = Up.patch().faceCells();
label start = Up.patch().start();
forAll(fc, i)
{
label facei = start+i;
label celli = fc[i];
label ownType = cellTypes[celli];
label neiType = neiCellTypes[facei-mesh.nInternalFaces()];
bool ownCalc =
(ownType == cellCellStencil::CALCULATED)
&& (neiType == cellCellStencil::INTERPOLATED);
if (ownCalc)
{
// Calculate flux w.r.t. calculated cell
scalar flux = phip[i];
if (flux < 0.0)
{
massIn[zoneID[celli]] -= flux;
}
else
{
adjustableMassOut[zoneID[celli]] += flux;
}
}
}
}
}The correction factor for each cell zone is calculated by the following line:
massCorr[zonei] = massIn[zonei]/adjustableMassOut[zonei];
. It is the ratio between the mass flux going over the boarder of a interpolated cell into a calculated cell to the mass flux leaving the calculated cell into an interpolated cell.
The correction factor massCorr is applied in the final step of the code. Here again a loop over all internal and also boundary faces is performed. The purpose of the correction factors is to adjust the global ratio over a mesh zone of mass flux from interpolated cells into a calculated cells to the mass flux from a calculated cells into an interpolated cells to one.
// Pass2: adjust fluxes
forAll(own, facei)
{
label zonei = zoneID[own[facei]]; // note:own and nei in same zone
label ownType = cellTypes[own[facei]];
label neiType = cellTypes[nei[facei]];
bool ownCalc =
(ownType == cellCellStencil::CALCULATED)
&& (neiType == cellCellStencil::INTERPOLATED);
bool neiCalc =
(ownType == cellCellStencil::INTERPOLATED)
&& (neiType == cellCellStencil::CALCULATED);
if (ownCalc || neiCalc)
{
// Calculate flux w.r.t. calculated cell
scalar flux = phi[facei];
if (neiCalc)
{
flux = -flux;
}
if (flux < 0.0)
{
phi[facei] /= Foam::sqrt(massCorr[zonei]);
}
else
{
phi[facei] *= Foam::sqrt(massCorr[zonei]);
}
}
}
forAll(bphi, patchi)
{
const fvPatchVectorField& Up = U.boundaryField()[patchi];
fvsPatchScalarField& phip = bphi[patchi];
const labelUList& fc = Up.patch().faceCells();
label start = Up.patch().start();
forAll(fc, i)
{
label facei = start+i;
label celli = fc[i];
label zonei = zoneID[celli]; // note:own and nei in same zone
label ownType = cellTypes[celli];
label neiType = neiCellTypes[facei-mesh.nInternalFaces()];
bool ownCalc =
(ownType == cellCellStencil::CALCULATED)
&& (neiType == cellCellStencil::INTERPOLATED);
bool neiCalc =
(ownType == cellCellStencil::INTERPOLATED)
&& (neiType == cellCellStencil::CALCULATED);
if (ownCalc || neiCalc)
{
// Calculate flux w.r.t. calculated cell
scalar flux = phip[i];
if (neiCalc)
{
flux = -flux;
}
if (flux < 0.0)
{
phip[i] /= Foam::sqrt(massCorr[zonei]);
}
else
{
phip[i] *= Foam::sqrt(massCorr[zonei]);
}
}
}
}
.
3 Overset interpolation
In this section we will analyze in more detail how the overset interpolation in OpenFOAM is achieved. Generally OpenFOAM uses an implicit interpolation. An overset interpolation can be written as:
-
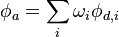 | (1) |
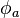 is the value of the variable
is the value of the variable  interpolated at the acceptor cell,
interpolated at the acceptor cell, 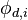 are the values of the donor cells and
are the values of the donor cells and 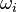 are the weights with which the values of the donor cells
are the weights with which the values of the donor cells 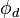 are applied. In an overset mesh, donor and acceptor cells are on disconnected mesh zones. This means that they are not connected over a face and the standard connectivity between cells cannot be used to construct the matrix. In brief, the procedure how to achieve an overset interpolation in OpenFOAM is the following: 1) Extend the existing ldu adressing and 2) insert the coefficients resulting from the overset interpolation in the matrix and 3) solve the linear system.
are applied. In an overset mesh, donor and acceptor cells are on disconnected mesh zones. This means that they are not connected over a face and the standard connectivity between cells cannot be used to construct the matrix. In brief, the procedure how to achieve an overset interpolation in OpenFOAM is the following: 1) Extend the existing ldu adressing and 2) insert the coefficients resulting from the overset interpolation in the matrix and 3) solve the linear system.
A brief description of how a matrix is constructed in OpenFOAM using a finite volume discretization can be found here https://openfoamwiki.net/index.php/OpenFOAM_guide/Matrices_in_OpenFOAM.
3.1 Computing the weights
In this section we will see how the weights for the overset interpolation are calculated. As example we will examine the inverse distance and the cell volume weight interpolation.
3.1.1 Inverse distance
For the inverse distance method the cellcellStencil is created in the function createStecil. The creation of the weight is also done here. The method in essence looks for each interpolated cell in meshZone A the closest cell in meshZone B. This is the first donor cell. After that, all face neighbors of the first donor cell are also taken as donor. This means that the cell stencil is very compact and second order accuracy can be achieved as best. Before we start with a short description of how the stencil is created, the code is shortly explained, who the closest donor to an interpolated cell is found. This task is performed in the function markDonor:
void Foam::cellCellStencils::inverseDistance::markDonors ( const globalIndex& globalCells, PstreamBuffers& pBufs, const PtrList<fvMeshSubset>& meshParts, const List<treeBoundBoxList>& meshBb, const labelList& allCellTypes, const label srcI, const label tgtI, labelListList& allStencil, labelList& allDonor ) const { const treeBoundBoxList& srcBbs = meshBb[srcI]; const treeBoundBoxList& tgtBbs = meshBb[tgtI]; const fvMesh& srcMesh = meshParts[srcI].subMesh(); const labelList& srcCellMap = meshParts[srcI].cellMap(); const fvMesh& tgtMesh = meshParts[tgtI].subMesh(); const pointField& tgtCc = tgtMesh.cellCentres(); const labelList& tgtCellMap = meshParts[tgtI].cellMap(); // 1. do processor-local src/tgt overlap { labelList tgtToSrcAddr; waveMethod::calculate(tgtMesh, srcMesh, tgtToSrcAddr); forAll(tgtCellMap, tgtCelli) { label srcCelli = tgtToSrcAddr[tgtCelli]; if (srcCelli != -1 && allCellTypes[srcCellMap[srcCelli]] != HOLE) { label celli = tgtCellMap[tgtCelli]; // TBD: check for multiple donors. Maybe better one? For // now check 'nearer' mesh if (betterDonor(tgtI, allDonor[celli], srcI)) { label globalDonor = globalCells.toGlobal(srcCellMap[srcCelli]); allStencil[celli].setSize(1); allStencil[celli][0] = globalDonor; allDonor[celli] = srcI; } } } } // 2. Send over tgtMesh bits that overlap src and do calculation on // srcMesh. // (remote) processors where the tgt overlaps my src DynamicList<label> tgtOverlapProcs(Pstream::nProcs()); // (remote) processors where the src overlaps my tgt DynamicList<label> srcOverlapProcs(Pstream::nProcs()); for (const int procI : Pstream::allProcs()) { if (procI != Pstream::myProcNo()) { if (tgtBbs[procI].overlaps(srcBbs[Pstream::myProcNo()])) { tgtOverlapProcs.append(procI); } if (srcBbs[procI].overlaps(tgtBbs[Pstream::myProcNo()])) { srcOverlapProcs.append(procI); } } } // Indices of tgtcells to send over to each processor List<DynamicList<label>> tgtSendCells(Pstream::nProcs()); forAll(srcOverlapProcs, i) { label procI = srcOverlapProcs[i]; tgtSendCells[procI].reserve(tgtMesh.nCells()/srcOverlapProcs.size()); } forAll(tgtCellMap, tgtCelli) { label celli = tgtCellMap[tgtCelli]; if (srcOverlapProcs.size()) { treeBoundBox subBb(cellBb(mesh_, celli)); subBb.min() -= smallVec_; subBb.max() += smallVec_; forAll(srcOverlapProcs, i) { label procI = srcOverlapProcs[i]; if (subBb.overlaps(srcBbs[procI])) { tgtSendCells[procI].append(tgtCelli); } } } } // Send target cell centres to overlapping processors pBufs.clear(); forAll(srcOverlapProcs, i) { label procI = srcOverlapProcs[i]; const labelList& cellIDs = tgtSendCells[procI]; UOPstream os(procI, pBufs); os << UIndirectList<point>(tgtCc, cellIDs); } pBufs.finishedSends(); // Receive bits of target processors; find; send back (void)srcMesh.tetBasePtIs(); forAll(tgtOverlapProcs, i) { label procI = tgtOverlapProcs[i]; UIPstream is(procI, pBufs); pointList samples(is); labelList donors(samples.size(), -1); forAll(samples, sampleI) { const point& sample = samples[sampleI]; label srcCelli = srcMesh.findCell(sample, polyMesh::CELL_TETS); if (srcCelli != -1 && allCellTypes[srcCellMap[srcCelli]] != HOLE) { donors[sampleI] = globalCells.toGlobal(srcCellMap[srcCelli]); } } // Use same pStreamBuffers to send back. UOPstream os(procI, pBufs); os << donors; } pBufs.finishedSends(); forAll(srcOverlapProcs, i) { label procI = srcOverlapProcs[i]; const labelList& cellIDs = tgtSendCells[procI]; UIPstream is(procI, pBufs); labelList donors(is); if (donors.size() != cellIDs.size()) { FatalErrorInFunction<< "problem : cellIDs:" << cellIDs.size() << " donors:" << donors.size() << abort(FatalError); } forAll(donors, donorI) { label globalDonor = donors[donorI]; if (globalDonor != -1) { label celli = tgtCellMap[cellIDs[donorI]]; // TBD: check for multiple donors. Maybe better one? For // now check 'nearer' mesh if (betterDonor(tgtI, allDonor[celli], srcI)) { allStencil[celli].setSize(1); allStencil[celli][0] = globalDonor; allDonor[celli] = srcI; } } } } }
Let's analyze shortly the above code. It is also quite instructive to analyze this part of code, to shortly highlight the method used to find the best donor over multiple overlapping regions. This regions can also be located on different processors. This code snippet is also used in order to have a short introduction of commutations between processors and the difficulties associated with it. When writing parallel code, one has always to keep in main, that the processors know only their own mesh if a domain decomposition is used. The code seen on the screen is executed by each processor without knowing anything about the other processor. In order to exchange information between processors an mpi communication has to be called. The first part of the code computes a one to one mapping between the source and target cell. This means that for each cell in the target mesh a cell is looked for which overlaps which th source cell. This is performed in the function
waveMethod::calculate(tgtMesh, srcMesh, tgtToSrcAddr);.
After that a source cell id is provided for each target cell in the same processor. This is done in the line
allStencil[celli][0] = globalDonor;
It is not said that all overlapping parts of a meshZone A with meshZone B are lying on the same processor. Now we have to check if a meshZone A lying in processor I is overalapping with a meshZone B lying in processor J. The first step to achieve this checking if the bounding boxes of meshZone A lying on the processor I where the code is executed overlaps with the mesh bounding box of meshZone B lying all other processors J:
// (remote) processors where the tgt overlaps my src
DynamicList<label> tgtOverlapProcs(Pstream::nProcs());
// (remote) processors where the src overlaps my tgt
DynamicList<label> srcOverlapProcs(Pstream::nProcs());
for (const int procI : Pstream::allProcs())
{
if (procI != Pstream::myProcNo())
{
if (tgtBbs[procI].overlaps(srcBbs[Pstream::myProcNo()]))
{
tgtOverlapProcs.append(procI);
}
if (srcBbs[procI].overlaps(tgtBbs[Pstream::myProcNo()]))
{
srcOverlapProcs.append(procI);
}
}
}
Once one knows which mesh part of meshZone A lying in processor I overalaps with a mesh part of meshZone B lying in processor J one can proceed to look for the cell which are closest to each other. At this stage the code executed in processor I does not know anything of the mesh in processor J. I order that processor I gets information about the mesh laying in processor J these information have to be sent over. In order to proceed further, the next step is to loop over each sell in meshZone A lying on the processor I where the code is executed and check if the cells overlap with the bounding box of meshZone B lying on another processor J. If is the case the cellId is stored in a listList with the first entry in the listList beeing the processor number J:
forAll(tgtCellMap, tgtCelli)
{
label celli = tgtCellMap[tgtCelli];
if (srcOverlapProcs.size())
{
treeBoundBox subBb(cellBb(mesh_, celli));
subBb.min() -= smallVec_;
subBb.max() += smallVec_;
forAll(srcOverlapProcs, i)
{
label procI = srcOverlapProcs[i];
if (subBb.overlaps(srcBbs[procI]))
{
tgtSendCells[procI].append(tgtCelli);
}
}
}
}
Having determined the cells lying in meshZone A on processor I where the code is executed which are overlapping with the bounding box of meshZone B lying on processor J, this infromation has to be sent to the corresponing processor J. We have to keep in mind, that at this stage processor I still does not know any informaiton of processo J:
forAll(srcOverlapProcs, i)
{
label procI = srcOverlapProcs[i];
const labelList& cellIDs = tgtSendCells[procI];
UOPstream os(procI, pBufs);
os << UIndirectList<point>(tgtCc, cellIDs);
}
pBufs.finishedSends();
The sending is done with the class UOPstream (see https://www.openfoam.com/documentation/guides/latest/api/UOPstream_8H_source.html) and it is waited until all processor have finished to send the data to the correspoing other processors (see the function finishedSend in https://www.openfoam.com/documentation/guides/latest/api/classFoam_1_1PstreamBuffers.html)..
The information of the cells which is sent over from processor I to processor J has to be recieved and processed. The processing steps is to determine for each sell of meshZone A lying on processor I the closest cell lying on processor J in cellzone B. Finally this information is sent back to processor I:
// Receive bits of target processors; find; send back
(void)srcMesh.tetBasePtIs();
forAll(tgtOverlapProcs, i)
{
label procI = tgtOverlapProcs[i];
UIPstream is(procI, pBufs);
pointList samples(is);
labelList donors(samples.size(), -1);
forAll(samples, sampleI)
{
const point& sample = samples[sampleI];
label srcCelli = srcMesh.findCell(sample, polyMesh::CELL_TETS);
if (srcCelli != -1 && allCellTypes[srcCellMap[srcCelli]] != HOLE)
{
donors[sampleI] = globalCells.toGlobal(srcCellMap[srcCelli]);
}
}
// Use same pStreamBuffers to send back.
UOPstream os(procI, pBufs);
os << donors;
}
pBufs.finishedSends();
The processor I has to recieve the date of processor J. The information which processor I recieves from processor J are the list global indeces of the cells in meshZone B lying closest to the cells in meshZone A on processor I. The information is stored in the listList
allStencil[celli][0] = globalDonor
forAll(srcOverlapProcs, i)
{
label procI = srcOverlapProcs[i];
const labelList& cellIDs = tgtSendCells[procI];
UOPstream os(procI, pBufs);
os << UIndirectList<point>(tgtCc, cellIDs);
}
pBufs.finishedSends();
</cpp>
forAll(srcOverlapProcs, i)
{
label procI = srcOverlapProcs[i];
const labelList& cellIDs = tgtSendCells[procI];
UIPstream is(procI, pBufs);
labelList donors(is);
if (donors.size() != cellIDs.size())
{
FatalErrorInFunction<< "problem : cellIDs:" << cellIDs.size()
<< " donors:" << donors.size() << abort(FatalError);
}
forAll(donors, donorI)
{
label globalDonor = donors[donorI];
if (globalDonor != -1)
{
label celli = tgtCellMap[cellIDs[donorI]];
// TBD: check for multiple donors. Maybe better one? For
// now check 'nearer' mesh
if (betterDonor(tgtI, allDonor[celli], srcI))
{
allStencil[celli].setSize(1);
allStencil[celli][0] = globalDonor;
allDonor[celli] = srcI;
}
}
}
void Foam::cellCellStencils::inverseDistance::createStencil ( const globalIndex& globalCells ) { // Send cell centre back to donor // ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~ // The complication is that multiple acceptors need the same donor // (but with different weights obviously) // So we do multi-pass: // - send over cc of acceptor for which we want stencil. // Consistently choose the acceptor with smallest magSqr in case of // multiple acceptors for the containing cell/donor. // - find the cell-cells and weights for the donor // - send back together with the acceptor cc // - use the acceptor cc to see if it was 'me' that sent it. If so // mark me as complete so it doesn't get involved in the next loop. // - loop until all resolved. // Special value for unused points const vector greatPoint(GREAT, GREAT, GREAT); boolList isValidDonor(mesh_.nCells(), true); forAll(cellTypes_, celli) { if (cellTypes_[celli] == HOLE) { isValidDonor[celli] = false; } } // Has acceptor been handled already? bitSet doneAcceptor(interpolationCells_.size()); while (true) { pointField samples(cellInterpolationMap().constructSize(), greatPoint); // Fill remote slots (override old content). We'll find out later // on which one has won and mark this one in doneAcceptor. label nSamples = 0; forAll(interpolationCells_, i) { if (!doneAcceptor[i]) { label cellI = interpolationCells_[i]; const point& cc = mesh_.cellCentres()[cellI]; const labelList& slots = cellStencil_[cellI]; if (slots.size() != 1) { FatalErrorInFunction<< "Problem:" << slots << abort(FatalError); } forAll(slots, slotI) { label elemI = slots[slotI]; //Pout<< " acceptor:" << cellI // << " at:" << mesh_.cellCentres()[cellI] // << " global:" << globalCells.toGlobal(cellI) // << " found in donor:" << elemI << endl; minMagSqrEqOp<point>()(samples[elemI], cc); } nSamples++; } } if (returnReduce(nSamples, sumOp<label>()) == 0) { break; } // Send back to donor. Make sure valid point takes priority mapDistributeBase::distribute<point, minMagSqrEqOp<point>, flipOp> ( Pstream::commsTypes::nonBlocking, List<labelPair>(), mesh_.nCells(), cellInterpolationMap().constructMap(), false, cellInterpolationMap().subMap(), false, samples, greatPoint, // nullValue minMagSqrEqOp<point>(), flipOp(), // negateOp UPstream::msgType(), cellInterpolationMap().comm() ); // All the donor cells will now have a valid cell centre. Construct a // stencil for these. DynamicList<label> donorCells(mesh_.nCells()); forAll(samples, cellI) { if (samples[cellI] != greatPoint) { donorCells.append(cellI); } } // Get neighbours (global cell and centre) of donorCells. labelListList donorCellCells(mesh_.nCells()); pointListList donorCellCentres(mesh_.nCells()); globalCellCells ( globalCells, mesh_, isValidDonor, donorCells, donorCellCells, donorCellCentres ); // Determine the weights. scalarListList donorWeights(mesh_.nCells()); forAll(donorCells, i) { label cellI = donorCells[i]; const pointList& donorCentres = donorCellCentres[cellI]; stencilWeights ( samples[cellI], donorCentres, donorWeights[cellI] ); } // Transfer the information back to the acceptor: // - donorCellCells : stencil (with first element the original donor) // - donorWeights : weights for donorCellCells cellInterpolationMap().distribute(donorCellCells); cellInterpolationMap().distribute(donorWeights); cellInterpolationMap().distribute(samples); // Check which acceptor has won and transfer forAll(interpolationCells_, i) { if (!doneAcceptor[i]) { label cellI = interpolationCells_[i]; const labelList& slots = cellStencil_[cellI]; if (slots.size() != 1) { FatalErrorInFunction << "Problem:" << slots << abort(FatalError); } label slotI = slots[0]; // Important: check if the stencil is actually for this cell if (samples[slotI] == mesh_.cellCentres()[cellI]) { cellStencil_[cellI].transfer(donorCellCells[slotI]); cellInterpolationWeights_[cellI].transfer ( donorWeights[slotI] ); // Mark cell as being done so it does not get sent over // again. doneAcceptor.set(i); } } } } // Re-do the mapDistribute List<Map<label>> compactMap; cellInterpolationMap_.reset ( new mapDistribute ( globalCells, cellStencil_, compactMap ) ); }
void Foam::cellCellStencil::globalCellCells ( const globalIndex& gi, const polyMesh& mesh, const boolList& isValidCell, const labelList& selectedCells, labelListList& cellCells, pointListList& cellCellCentres ) { // For selected cells determine the face neighbours (in global numbering) const pointField& cellCentres = mesh.cellCentres(); const labelList& faceOwner = mesh.faceOwner(); const labelList& faceNeighbour = mesh.faceNeighbour(); const cellList& cells = mesh.cells(); // 1. Determine global cell number on other side of coupled patches labelList globalCellIDs(identity(gi.localSize(), gi.localStart())); labelList nbrGlobalCellIDs; syncTools::swapBoundaryCellList ( mesh, globalCellIDs, nbrGlobalCellIDs ); pointField nbrCellCentres; syncTools::swapBoundaryCellList ( mesh, cellCentres, nbrCellCentres ); boolList nbrIsValidCell; syncTools::swapBoundaryCellList ( mesh, isValidCell, nbrIsValidCell ); // 2. Collect cell and all its neighbours cellCells.setSize(mesh.nCells()); cellCellCentres.setSize(cellCells.size()); forAll(selectedCells, i) { label celli = selectedCells[i]; const cell& cFaces = cells[celli]; labelList& stencil = cellCells[celli]; pointList& stencilPoints = cellCellCentres[celli]; stencil.setSize(cFaces.size()+1); stencilPoints.setSize(stencil.size()); label compacti = 0; // First entry is cell itself if (isValidCell[celli]) { stencil[compacti] = globalCellIDs[celli]; stencilPoints[compacti++] = cellCentres[celli]; } // Other entries are cell neighbours forAll(cFaces, i) { label facei = cFaces[i]; label bFacei = facei-mesh.nInternalFaces(); label own = faceOwner[facei]; label nbrCelli; point nbrCc; bool isValid = false; if (bFacei >= 0) { nbrCelli = nbrGlobalCellIDs[bFacei]; nbrCc = nbrCellCentres[bFacei]; isValid = nbrIsValidCell[bFacei]; } else { if (own != celli) { nbrCelli = gi.toGlobal(own); nbrCc = cellCentres[own]; isValid = isValidCell[own]; } else { label nei = faceNeighbour[facei]; nbrCelli = gi.toGlobal(nei); nbrCc = cellCentres[nei]; isValid = isValidCell[nei]; } } if (isValid) { SubList<label> current(stencil, compacti); if (!current.found(nbrCelli)) { stencil[compacti] = nbrCelli; stencilPoints[compacti++] = nbrCc; } } } stencil.setSize(compacti); stencilPoints.setSize(compacti); } }
void Foam::cellCellStencils::inverseDistance::stencilWeights ( const point& sample, const pointList& donorCcs, scalarList& weights ) const { // Inverse-distance weighting weights.setSize(donorCcs.size()); scalar sum = 0.0; forAll(donorCcs, i) { scalar d = mag(sample-donorCcs[i]); if (d > ROOTVSMALL) { weights[i] = 1.0/d; sum += weights[i]; } else { // Short circuit weights = 0.0; weights[i] = 1.0; return; } } forAll(weights, i) { weights[i] /= sum; } }
3.1.2 Cell volume weight
3.2 Extended ldu addressing
Let's start to analyze how the ldu addressing in OpenFOAM is extended by the additional connectivity resulting from the overset interpolation. As first step let's recap how the connectivity between cells and the entries in the matrix in OpenFOAM are stored. The connectivity between the cells are stored in two arrays with the same length:
lowerAddr() upperAddr()
In the
lowerAddr()
array the smaller index of the two coupled cells are stored and in the
upperAddr()
array the higher index of two coupled cells is stored. In OpenFOAM this both arrays have the length equal to the number of faces since only adjacent cells are used for the discretization.
The corresponding entries in the matrix are stored in other two arrays:
lower() upper()
In order to account for the additional coupling between cell centers introduced by the overset meh approach, new entries in the matrix have to be accomplished. This means that the length of the above four arrays has to be enlarged. In the arrays
lowerAddr() upperAddr()
the additional indexes of the cells coupled by the overset interpolation has to be added. In the arrays
lower()
upper()
the coefficients in the matrix resulting from the coupling have to be added. This is done during step when the mesh is updated:
bool changed = mesh.update();
In case of solvers which use the overset method, the mesh instance is of type dynamicOversetFvMesh. The method update can be found in dynamicOversetFvMesh.C:
bool Foam::dynamicOversetFvMesh::update() { //if (dynamicMotionSolverFvMesh::update()) if (dynamicMotionSolverListFvMesh::update()) { // Calculate the local extra faces for the interpolation. Note: could // let demand-driven lduAddr() trigger it but just to make sure. updateAddressing(); // Addressing and/or weights have changed. Make interpolated cells // up to date with donors interpolateFields(); return true; } return false; }
The updating of the addressing happens function fvMeshPrimitiveLduAddressing::addAddressing in the file fvMeshPrimitiveLduAddressing.C:
Foam::labelList Foam::fvMeshPrimitiveLduAddressing::addAddressing ( const lduAddressing& addr, const labelListList& nbrCells, label& nExtraFaces, labelList& newLowerAddr, labelList& newUpperAddr, labelListList& nbrCellFaces, const globalIndex& globalNumbering, const labelList& globalCellIDs, labelListList& localFaceCells, labelListList& remoteFaceCells ) { label nCells = addr.size(); label nFaces = addr.upperAddr().size(); labelList nProcFaces(Pstream::nProcs(), Zero); // Count additional faces nExtraFaces = 0; forAll(nbrCells, cellI) { const labelList& nbrs = nbrCells[cellI]; forAll(nbrs, nbrI) { if (nbrs[nbrI] < nCells) { // Local cell if (triIndex(addr, cellI, nbrs[nbrI]) == -1) { nExtraFaces++; } } else { label globalNbr = globalCellIDs[nbrs[nbrI]]; label procI = globalNumbering.whichProcID(globalNbr); nProcFaces[procI]++; } } } // Create space for extra addressing newLowerAddr.setSize(nFaces + nExtraFaces); newUpperAddr.setSize(nFaces + nExtraFaces); // Copy existing addressing SubList<label>(newLowerAddr, nFaces) = addr.lowerAddr(); SubList<label>(newUpperAddr, nFaces) = addr.upperAddr(); // Per processor its local cells we want localFaceCells.setSize(Pstream::nProcs()); remoteFaceCells.setSize(Pstream::nProcs()); forAll(nProcFaces, procI) { localFaceCells[procI].setSize(nProcFaces[procI]); remoteFaceCells[procI].setSize(nProcFaces[procI]); } nProcFaces = 0; nbrCellFaces.setSize(nbrCells.size()); forAll(nbrCells, cellI) { const labelList& nbrs = nbrCells[cellI]; labelList& faces = nbrCellFaces[cellI]; faces.setSize(nbrs.size()); forAll(nbrs, nbrI) { label nbrCellI = nbrs[nbrI]; if (nbrCellI < nCells) { // Find neighbour cell in owner list label faceI = triIndex(addr, cellI, nbrCellI); if (faceI == -1) { faceI = nFaces++; newLowerAddr[faceI] = min(cellI, nbrCellI); newUpperAddr[faceI] = max(cellI, nbrCellI); } faces[nbrI] = faceI; } else { // Remote cell faces[nbrI] = -1; label globalNbr = globalCellIDs[nbrCellI]; label procI = globalNumbering.whichProcID(globalNbr); label remoteCellI = globalNumbering.toLocal(procI, globalNbr); label procFaceI = nProcFaces[procI]++; localFaceCells[procI][procFaceI] = cellI; remoteFaceCells[procI][procFaceI] = remoteCellI; } } } // Reorder upper-triangular labelList oldToNew ( lduPrimitiveMesh::upperTriOrder ( addr.size(), newLowerAddr, newUpperAddr ) ); // Shuffle face-to-cell addressing inplaceReorder(oldToNew, newLowerAddr); inplaceReorder(oldToNew, newUpperAddr); // Update cell-to-face addressing forAll(nbrCellFaces, cellI) { inplaceRenumber(oldToNew, nbrCellFaces[cellI]); } return oldToNew; }
The information of the indices of the coupled cells is contained in the cellcellStencil object. It is basically a list of lists. For each index of the interpolated cells it contains the indices of the donor cell. If one knows the pairing index donor cell and acceptor cell, one can update the lduaddrissing. The lower index can be inserted in the lowerAddr array and the higer index can be inserted in the upperAdrr array. This is down by the following code snippet:
if (nbrCellI < nCells)
{
// Find neighbour cell in owner list
label faceI = triIndex(addr, cellI, nbrCellI);
if (faceI == -1)
{
faceI = nFaces++;
newLowerAddr[faceI] = min(cellI, nbrCellI);
newUpperAddr[faceI] = max(cellI, nbrCellI);
}
faces[nbrI] = faceI;
}3.3 Modification of the ldu matrix
After having extended the addressing by the additional connectivity resulting form the overset interpolation, the coefficient resulting from the overset interpolation have to be inserted in the matrix and the matrix have to be solve. The function call very the matrix is solved happens in dynamicOversetFvMesh::solve. The function takes the matrix resulting from the standard finite volume discrcretization (done in the files Ueqn.H, peqn.H a.s.o), adds the coefficients resulting from the overset interpolation and solves the matrix:
template<class Type> Foam::SolverPerformance<Type> Foam::dynamicOversetFvMesh::solve ( fvMatrix<Type>& m, const dictionary& dict ) const { typedef GeometricField<Type, fvPatchField, volMesh> GeoField; // Check we're running with bcs that can handle implicit matrix manipulation typename GeoField::Boundary& bpsi = const_cast<GeoField&>(m.psi()).boundaryFieldRef(); bool hasOverset = false; forAll(bpsi, patchi) { if (isA<oversetFvPatchField<Type>>(bpsi[patchi])) { hasOverset = true; break; } } if (!hasOverset) { if (debug) { Pout<< "dynamicOversetFvMesh::solve() :" << " bypassing matrix adjustment for field " << m.psi().name() << endl; } //return dynamicMotionSolverFvMesh::solve(m, dict); return dynamicFvMesh::solve(m, dict); } if (debug) { Pout<< "dynamicOversetFvMesh::solve() :" << " adjusting matrix for interpolation for field " << m.psi().name() << endl; } // Calculate stabilised diagonal as normalisation for interpolation const scalarField norm(normalisation(m)); if (debug) { volScalarField scale ( IOobject ( m.psi().name() + "_normalisation", this->time().timeName(), *this, IOobject::NO_READ, IOobject::NO_WRITE, false ), *this, dimensionedScalar(dimless, Zero) ); scale.ref().field() = norm; correctBoundaryConditions < volScalarField, oversetFvPatchField<scalar> >(scale.boundaryFieldRef(), false); scale.write(); if (debug) { Pout<< "dynamicOversetFvMesh::solve() :" << " writing matrix normalisation for field " << m.psi().name() << " to " << scale.name() << endl; } } // Switch to extended addressing (requires mesh::update() having been // called) active(true); // Adapt matrix scalarField oldUpper(m.upper()); scalarField oldLower(m.lower()); FieldField<Field, Type> oldInt(m.internalCoeffs()); FieldField<Field, Type> oldBou(m.boundaryCoeffs()); const label oldSize = bpsi.size(); addInterpolation(m, norm); // Swap psi values so added patches have patchNeighbourField correctBoundaryConditions<GeoField, calculatedProcessorFvPatchField<Type>> ( bpsi, true ); // Print a bit //write(Pout, m, lduAddr(), interfaces()); //{ // const fvSolution& sol = static_cast<const fvSolution&>(*this); // const dictionary& pDict = sol.subDict("solvers").subDict("p"); // writeAgglomeration(GAMGAgglomeration::New(m, pDict)); //} // Use lower level solver //SolverPerformance<Type> s(dynamicMotionSolverFvMesh::solve(m, dict)); SolverPerformance<Type> s(dynamicFvMesh::solve(m, dict)); // Restore boundary field bpsi.setSize(oldSize); // Restore matrix m.upper().transfer(oldUpper); m.lower().transfer(oldLower); m.internalCoeffs().transfer(oldInt); m.boundaryCoeffs().transfer(oldBou); // Switch to original addressing active(false); return s; }
The actual interpolation is done in the function dynamicOversetFvMesh::addInterpolation:
template<class Type> void Foam::dynamicOversetFvMesh::addInterpolation ( fvMatrix<Type>& m, const scalarField& normalisation ) const { const cellCellStencilObject& overlap = Stencil::New(*this); const List<scalarList>& wghts = overlap.cellInterpolationWeights(); const labelListList& stencil = overlap.cellStencil(); const labelList& cellIDs = overlap.interpolationCells(); const scalarList& factor = overlap.cellInterpolationWeight(); const labelUList& types = overlap.cellTypes(); // Force asymmetric matrix (if it wasn't already) scalarField& lower = m.lower(); scalarField& upper = m.upper(); Field<Type>& source = m.source(); scalarField& diag = m.diag(); // Get the addressing. Note that the addressing is now extended with // any interpolation faces. const lduAddressing& addr = lduAddr(); const labelUList& upperAddr = addr.upperAddr(); const labelUList& lowerAddr = addr.lowerAddr(); const labelUList& ownerStartAddr = addr.ownerStartAddr(); const labelUList& losortAddr = addr.losortAddr(); const lduInterfacePtrsList& interfaces = allInterfaces_; if (!isA<fvMeshPrimitiveLduAddressing>(addr)) { FatalErrorInFunction << "Problem : addressing is not fvMeshPrimitiveLduAddressing" << exit(FatalError); } // 1. Adapt lduMatrix for additional faces and new ordering upper.setSize(upperAddr.size(), 0.0); inplaceReorder(reverseFaceMap_, upper); lower.setSize(lowerAddr.size(), 0.0); inplaceReorder(reverseFaceMap_, lower); //const label nOldInterfaces = dynamicMotionSolverFvMesh::interfaces().size(); const label nOldInterfaces = dynamicFvMesh::interfaces().size(); if (interfaces.size() > nOldInterfaces) { // Extend matrix coefficients m.internalCoeffs().setSize(interfaces.size()); m.boundaryCoeffs().setSize(interfaces.size()); // 1b. Adapt for additional interfaces for ( label patchi = nOldInterfaces; patchi < interfaces.size(); patchi++ ) { const labelUList& fc = interfaces[patchi].faceCells(); m.internalCoeffs().set(patchi, new Field<Type>(fc.size(), Zero)); m.boundaryCoeffs().set(patchi, new Field<Type>(fc.size(), Zero)); } // 1c. Adapt field for additional interfaceFields (note: solver uses // GeometricField::scalarInterfaces() to get hold of interfaces) typedef GeometricField<Type, fvPatchField, volMesh> GeoField; typename GeoField::Boundary& bfld = const_cast<GeoField&>(m.psi()).boundaryFieldRef(); bfld.setSize(interfaces.size()); // This gets quite interesting: we do not want to add additional // fvPatches (since direct correspondence to polyMesh) so instead // add a reference to an existing processor patch label addPatchi = 0; for (label patchi = 0; patchi < nOldInterfaces; patchi++) { if (isA<processorFvPatch>(bfld[patchi].patch())) { addPatchi = patchi; break; } } for ( label patchi = nOldInterfaces; patchi < interfaces.size(); patchi++ ) { bfld.set ( patchi, new calculatedProcessorFvPatchField<Type> ( interfaces[patchi], bfld[addPatchi].patch(), // dummy processorFvPatch m.psi() ) ); } } // 2. Adapt fvMatrix level: faceFluxCorrectionPtr // Question: do we need to do this? // This seems to be set/used only by the gaussLaplacianScheme and // fvMatrix:correction, both of which are outside the linear solver. // Clear out existing connections on cells to be interpolated // ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~ // Note: could avoid doing the zeroing of the new faces since these // are set to zero anyway. forAll(upperAddr, facei) { if (types[upperAddr[facei]] == cellCellStencil::INTERPOLATED) { // Disconnect upper from lower label celli = upperAddr[facei]; lower[facei] *= 1.0-factor[celli]; } if (types[lowerAddr[facei]] == cellCellStencil::INTERPOLATED) { // Disconnect lower from upper label celli = lowerAddr[facei]; upper[facei] *= 1.0-factor[celli]; } } for (label patchi = 0; patchi < nOldInterfaces; ++patchi) { const labelUList& fc = addr.patchAddr(patchi); Field<Type>& intCoeffs = m.internalCoeffs()[patchi]; Field<Type>& bouCoeffs = m.boundaryCoeffs()[patchi]; forAll(fc, i) { label celli = fc[i]; { if (types[celli] == cellCellStencil::INTERPOLATED) { scalar f = factor[celli]; intCoeffs[i] *= 1.0-f; bouCoeffs[i] *= 1.0-f; } else if (types[celli] == cellCellStencil::HOLE) { intCoeffs[i] = pTraits<Type>::zero; bouCoeffs[i] = pTraits<Type>::zero; } } } } // Modify matrix // ~~~~~~~~~~~~~ // Do hole cells. Note: maybe put into interpolationCells() loop above? forAll(types, celli) { if (types[celli] == cellCellStencil::HOLE) { label startLabel = ownerStartAddr[celli]; label endLabel = ownerStartAddr[celli + 1]; for (label facei = startLabel; facei < endLabel; facei++) { upper[facei] = 0.0; } startLabel = addr.losortStartAddr()[celli]; endLabel = addr.losortStartAddr()[celli + 1]; for (label i = startLabel; i < endLabel; i++) { label facei = losortAddr[i]; lower[facei] = 0.0; } diag[celli] = normalisation[celli]; source[celli] = normalisation[celli]*m.psi()[celli]; } } forAll(cellIDs, i) { label celli = cellIDs[i]; const scalar f = factor[celli]; const scalarList& w = wghts[celli]; const labelList& nbrs = stencil[celli]; const labelList& nbrFaces = stencilFaces_[celli]; const labelList& nbrPatches = stencilPatches_[celli]; if (types[celli] == cellCellStencil::HOLE) { FatalErrorInFunction << "Found HOLE cell " << celli << " at:" << C()[celli] << " . Should this be in interpolationCells()????" << abort(FatalError); } else { // Create interpolation stencil diag[celli] *= (1.0-f); source[celli] *= (1.0-f); forAll(nbrs, nbri) { label patchi = nbrPatches[nbri]; label facei = nbrFaces[nbri]; if (patchi == -1) { label nbrCelli = nbrs[nbri]; // Add the coefficients const scalar s = normalisation[celli]*f*w[nbri]; scalar& u = upper[facei]; scalar& l = lower[facei]; if (celli < nbrCelli) { diag[celli] += s; u += -s; } else { diag[celli] += s; l += -s; } } else { const scalar s = normalisation[celli]*f*w[nbri]; m.boundaryCoeffs()[patchi][facei] += pTraits<Type>::one*s; m.internalCoeffs()[patchi][facei] += pTraits<Type>::one*s; // Note: do NOT add to diagonal - this is in the // internalCoeffs and gets added to the diagonal // inside fvMatrix::solve } } } } }
In order to understand what is done let's go step by step through the function. In the first part of the function the information about the cells involved in the overset interpolation and the weights used for the interpolation are retrieved:
const cellCellStencilObject& overlap = Stencil::New(*this);
const List<scalarList>& wghts = overlap.cellInterpolationWeights();
const labelListList& stencil = overlap.cellStencil();
const labelList& cellIDs = overlap.interpolationCells();
const scalarList& factor = overlap.cellInterpolationWeight();
const labelUList& types = overlap.cellTypes();overlap.cellInterpolationWeights();
returns a list of scalar lists where for each label of interpolated cells index the weights for the interpolation are stored.
overlap.cellStencil();
returns a list of labels where for each index interpolated cells the indeces of donor cells is stored.
overlap.interpolationCells()
returns the indeces of the interpolated cells
overlap.cellInterpolationWeight();
returns 1 for a interpolated cell and 0 for a calculated cell
overlap.cellTypes();
returns the type of cells (interpolated, hole or calculated).
As next point let's shortly recap how OpenFOAM stores the matrix and the indexes of the cells which are coupled in the matrix: lowerAddr stores the smaller index of a cell delimited by a face and upperAddr stores the higher index delimited by the same face. The entries in the two arrays are ordered in terms of face number. The coefficents of the matrix are stored in the arrays lower, upper and diag. diag contains the diagonal components of the matrix. lower() contains the coeffcients of the matrix M[upperAddr[facei]][lowerAddr[facei]] and upper() contain the matrix coefficients M[lowerAddr[facei]][upperAddr[facei]]. The entries of the source term (the right hand side vector) are stored in source.
The actual accounting of the overset interpolation at matrix level happens in two steps. The first step is to clear out the existing connections between the interpolated cells and its neighbors. This is has to be done since in the first step the usual matrix (without overset interpolation) is build using a standard finite volume discretization. This can be seen looking at the source code of e.g. Ueqn.H:
// Solve the Momentum equation MRF.correctBoundaryVelocity(U); tmp<fvVectorMatrix> tUEqn ( fvm::ddt(U) + fvm::div(phi, U) + MRF.DDt(U) + turbulence->divDevReff(U) == fvOptions(U) ); fvVectorMatrix& UEqn = tUEqn.ref(); UEqn.relax(); fvOptions.constrain(UEqn); if (pimple.momentumPredictor()) { solve(UEqn == -cellMask*fvc::grad(p)); fvOptions.correct(U); }
Here fist the matrix is build using standard finte volume operators (like ddt(U) ore div(phi,U)). At this stage it is not distinquished if the cells are CALCULATED, INTERPOLATED or HOLE. The actual overset interpolation at matrix level is done in the function solve. Since the standard fine volume operators like ddt(U) or div(phi,U) do not know anything about the overset interpolation, the connections between the interpolated cells and the surrounding cells have to be deleted. The connections between the donor and acceptor cells are already accounted in the extention of the ldu adressing. Clearing out the connections between the interpolated cells and its neighbors means that all off-diaganol coefficients of the matrix equation belonging to the interpolated cells are set to zero. That means if a cell with the lable n is an interpolated cells, all off diagonal elements of the nth row of th the original matrix have to be set to zero. The clearing out is done in the following code snipet:
forAll(upperAddr, facei)
{
if (types[upperAddr[facei]] == cellCellStencil::INTERPOLATED)
{
// Disconnect upper from lower
label celli = upperAddr[facei];
lower[facei] *= 1.0-factor[celli];
}
if (types[lowerAddr[facei]] == cellCellStencil::INTERPOLATED)
{
// Disconnect lower from upper
label celli = lowerAddr[facei];
upper[facei] *= 1.0-factor[celli];
}
}
for (label patchi = 0; patchi < nOldInterfaces; ++patchi)
{
const labelUList& fc = addr.patchAddr(patchi);
Field<Type>& intCoeffs = m.internalCoeffs()[patchi];
Field<Type>& bouCoeffs = m.boundaryCoeffs()[patchi];
forAll(fc, i)
{
label celli = fc[i];
{
if (types[celli] == cellCellStencil::INTERPOLATED)
{
scalar f = factor[celli];
intCoeffs[i] *= 1.0-f;
bouCoeffs[i] *= 1.0-f;
}
else if (types[celli] == cellCellStencil::HOLE)
{
intCoeffs[i] = pTraits<Type>::zero;
bouCoeffs[i] = pTraits<Type>::zero;
}
}
}
}
In the above code a loop over all faces is performed. First it is checked if the neighour cells of a face (the index of the neighbore cells are stored in the array upperAddr[] ) is of type INTERPOLATED. Remember that the array lower() stores the lower triangle entries of the matrix A according the following order A[upperAddr[facei]][lowerAddr[facei]]. Since we have to set the entries of the row of the matrix A to zero which give a contributio to the equation for the interpolated cell, lower[facei] has to be set to zero. An analogous operation is also performed for the owner cells. In this way the coupling between the interpolated cells and its neighbors coming from the finite volume descretization are irrased.
The next step performed is to acount for the overset interpolation at matrix level. As first step let's recall the relation between the donor and acceptor cells coming from the overset interpolatoin:
-
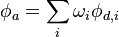 | (1) |
This equation establishes a linear relation between the interpoloted cells and the donor cells. Note that the coupling is only one way: The interpolated cells are influenced by the donor cells but not visa versa. The solution of the donor cells is achieved by standart finite volume discretization. The following code snipet shows how the overset interpolation is achived on matrix level:
forAll(cellIDs, i) { label celli = cellIDs[i]; const scalar f = factor[celli]; const scalarList& w = wghts[celli]; const labelList& nbrs = stencil[celli]; const labelList& nbrFaces = stencilFaces_[celli]; const labelList& nbrPatches = stencilPatches_[celli]; if (types[celli] == cellCellStencil::HOLE) { FatalErrorInFunction << "Found HOLE cell " << celli << " at:" << C()[celli] << " . Should this be in interpolationCells()????" << abort(FatalError); } else { // Create interpolation stencil diag[celli] *= (1.0-f); source[celli] *= (1.0-f); forAll(nbrs, nbri) { label patchi = nbrPatches[nbri]; label facei = nbrFaces[nbri]; if (patchi == -1) { label nbrCelli = nbrs[nbri]; // Add the coefficients const scalar s = normalisation[celli]*f*w[nbri]; scalar& u = upper[facei]; scalar& l = lower[facei]; if (celli < nbrCelli) { diag[celli] += s; u += -s; } else { diag[celli] += s; l += -s; } } else { // Patch face. Store in boundaryCoeffs. Note sign change. //const label globalCelli = globalCellIDs[nbrs[nbri]]; //const label proci = // globalNumbering.whichProcID(globalCelli); //const label remoteCelli = // globalNumbering.toLocal(proci, globalCelli); // //Pout<< "for cell:" << celli // << " need weight from remote slot:" << nbrs[nbri] // << " proc:" << proci << " remote cell:" << remoteCelli // << " patch:" << patchi // << " patchFace:" << facei // << " weight:" << w[nbri] // << endl; const scalar s = normalisation[celli]*f*w[nbri]; m.boundaryCoeffs()[patchi][facei] += pTraits<Type>::one*s; m.internalCoeffs()[patchi][facei] += pTraits<Type>::one*s; // Note: do NOT add to diagonal - this is in the // internalCoeffs and gets added to the diagonal // inside fvMatrix::solve } } } }
The above code starts with a loop over all interpolated cells. The loop starts by retrieving the information required for the interpolation: f is 1 for an interpolated cells and 0 else, the weights w required for the interpolation, the indexes nbrs of the donor cells for each interpolated cells. The list stencilFaces_[celli] containts the position of the donor cells for the cell with the index celli in the extended upperAddr and lowerAddr array and stencilPatches_[celli] contains the id of the neighbor processor if the donor cell lays on a processor boundary.
The loop continous by setting the diagonal and the source to zero. Note that the diagonal will be set to one in the later stage of the loop since the sum of the weights is one. After that the coefficients of the matrix resulting from the overset interpolation are written. The lable facei determines the position in the lower and upper array very the coupling coefficients between donor and acceptor have to be stored. As fininal step the entries in the lower or upper array are written depending on which of the donor or acceptor cell lable is higher.
4 Determination of the cell types
In this section we will see how the three different cell types are determined in OpenFOAM.
4.1 Hole cells
Hole cells in the frame work of an overset mesh are cells which are excluded from the calculation. The scope of the exclusion is to mimic the behavior of the solid body: Cells in the mesh inside the region delimited by the solid body should not have any influence on the solution of the region which are outside the region delimited by the body. The algorithm for the search of the hole cells works in two steps: 1) Find the cells in the mesh which are cut by wall patches 2) Find cells inside the region cut by the walls.
4.1.1 Find the cells in the mesh which are cut by wall patches
Unfortunately the algorithms used to find the cells in a cell zone which are cut by patches present in another zone are not the same for all method to calculate the cell cell stencil later used for the interpolation. For this reason the methods used in the inverse distance and the cell volume weight are presented briefly.
4.1.1.1 Inverse distance weight method
The first step of the algorithm to mark the hole cells at the boundaries is to mark the boundary cells in each cell Zone. The basic idea of the to decompose the mesh bounding box into voxels. A voxel represents a value in a regular 3D grid. It can be seen as a 3D analogous of a pixel. For an explanation see https://en.wikipedia.org/wiki/Voxel. The list of voxels for each cell zone are stored in patchParts. Later all loop over all boundary cells is performed. It is checked if the boundary cell bounding box overlaps with the mesh bounding box. If it overlaps, the voxels within the boundary cell bounding box are marked. This is done in the following code snippet.
forAll(patchParts, zoneI)
{
patchParts.set
(
zoneI,
new PackedList<2>
(
patchDivisions[zoneI][0]
*patchDivisions[zoneI][1]
*patchDivisions[zoneI][2]
)
);
markBoundaries
(
meshParts[zoneI].subMesh(),
smallVec_,
patchBb[zoneI][Pstream::myProcNo()],
patchDivisions[zoneI],
patchParts[zoneI],
meshParts[zoneI].cellMap(),
allPatchTypes
);
}The patchBb is the patch bounding box and is usually equal to the mesh bounding box:
{
treeBoundBox globalPatchBb;
if (dict_.readIfPresent("searchBox", globalPatchBb))
{
// All processors, all zones have the same bounding box
patchBb = treeBoundBoxList(Pstream::nProcs(), globalPatchBb);
}
else
{
// Use the meshBb (differing per zone, per processor)
patchBb = meshBb;
}
}The mark boundary function which marks the voxels at the boundary with the value PATCH or OVERSET looks at follows.
void Foam::cellCellStencils::inverseDistance::markBoundaries ( const fvMesh& mesh, const vector& smallVec, const boundBox& bb, const labelVector& nDivs, PackedList<2>& patchTypes, const labelList& cellMap, labelList& patchCellTypes ) { // Mark all voxels that overlap the bounding box of any patch const fvBoundaryMesh& pbm = mesh.boundary(); patchTypes = patchCellType::OTHER; // Mark wall boundaries forAll(pbm, patchI) { const fvPatch& fvp = pbm[patchI]; const labelList& fc = fvp.faceCells(); if (!fvPatch::constraintType(fvp.type())) { //Info<< "Marking cells on proper patch " << fvp.name() // << " with type " << fvp.type() << endl; const polyPatch& pp = fvp.patch(); forAll(pp, i) { // Mark in overall patch types patchCellTypes[cellMap[fc[i]]] = patchCellType::PATCH; // Mark in voxel mesh boundBox faceBb(pp.points(), pp[i], false); faceBb.min() -= smallVec; faceBb.max() += smallVec; if (bb.overlaps(faceBb)) { fill(patchTypes, bb, nDivs, faceBb, patchCellType::PATCH); } } } } // Override with overset boundaries forAll(pbm, patchI) { const fvPatch& fvp = pbm[patchI]; const labelList& fc = fvp.faceCells(); if (isA<oversetFvPatch>(fvp)) { //Info<< "Marking cells on overset patch " << fvp.name() << endl; const polyPatch& pp = fvp.patch(); forAll(pp, i) { // Mark in overall patch types patchCellTypes[cellMap[fc[i]]] = patchCellType::OVERSET; // Mark in voxel mesh boundBox faceBb(pp.points(), pp[i], false); faceBb.min() -= smallVec; faceBb.max() += smallVec; if (bb.overlaps(faceBb)) { fill(patchTypes, bb, nDivs, faceBb, patchCellType::OVERSET); } } } } }
The assigning of the voxels with the value PATCH or OVERSET is done in the following code snippet.
void Foam::cellCellStencils::inverseDistance::fill ( PackedList<2>& elems, const boundBox& bb, const labelVector& nDivs, const boundBox& subBb, const unsigned int val ) { labelVector minIds(index3(bb, nDivs, subBb.min())); labelVector maxIds(index3(bb, nDivs, subBb.max())); for (direction cmpt = 0; cmpt < 3; cmpt++) { if (maxIds[cmpt] < 0 || minIds[cmpt] > nDivs[cmpt]) { return; } } labelVector maxIndex(labelVector(nDivs[0]-1, nDivs[1]-1, nDivs[2]-1)); minIds = max(labelVector::zero, minIds); maxIds = min(maxIndex, maxIds); for (label i = minIds[0]; i <= maxIds[0]; i++) { for (label j = minIds[1]; j <= maxIds[1]; j++) { for (label k = minIds[2]; k <= maxIds[2]; k++) { label i1 = index(nDivs, labelVector(i, j, k)); elems[i1] = val; } } } }
The next step is to mark the cells in a mesh Zone A which are cut by a voxel in mesh Zone B marked as PATCH as HOLE. This is done here:
for (label srcI = 0; srcI < meshParts.size()-1; srcI++) { for (label tgtI = srcI+1; tgtI < meshParts.size(); tgtI++) { markPatchesAsHoles ( pBufs, meshParts, patchBb, patchDivisions, patchParts, srcI, tgtI, allCellTypes ); markPatchesAsHoles ( pBufs, meshParts, patchBb, patchDivisions, patchParts, tgtI, srcI, allCellTypes ); } }
The function markPatchesAsHoles looks as follows. The basic idea of the algorithm is loop over all cells of meshZone A and look if they cut any of the voxels defining the boundary of meshZone B. Around each cell of meshZone A a bounding box is build. The check for the interaction of two hexaedra is done very easily.
void Foam::cellCellStencils::inverseDistance::markPatchesAsHoles ( PstreamBuffers& pBufs, const PtrList<fvMeshSubset>& meshParts, const List<treeBoundBoxList>& patchBb, const List<labelVector>& patchDivisions, const PtrList<PackedList<2>>& patchParts, const label srcI, const label tgtI, labelList& allCellTypes ) const { const treeBoundBoxList& srcPatchBbs = patchBb[srcI]; const treeBoundBoxList& tgtPatchBbs = patchBb[tgtI]; const labelList& tgtCellMap = meshParts[tgtI].cellMap(); // 1. do processor-local src-tgt patch overlap { const treeBoundBox& srcPatchBb = srcPatchBbs[Pstream::myProcNo()]; const treeBoundBox& tgtPatchBb = tgtPatchBbs[Pstream::myProcNo()]; if (srcPatchBb.overlaps(tgtPatchBb)) { const PackedList<2>& srcPatchTypes = patchParts[srcI]; const labelVector& zoneDivs = patchDivisions[srcI]; forAll(tgtCellMap, tgtCelli) { label celli = tgtCellMap[tgtCelli]; treeBoundBox cBb(cellBb(mesh_, celli)); cBb.min() -= smallVec_; cBb.max() += smallVec_; if ( overlaps ( srcPatchBb, zoneDivs, srcPatchTypes, cBb, patchCellType::PATCH ) ) { allCellTypes[celli] = HOLE; } } } } // 2. Send over srcMesh bits that overlap tgt and do calculation pBufs.clear(); for (const int procI : Pstream::allProcs()) { if (procI != Pstream::myProcNo()) { const treeBoundBox& srcPatchBb = srcPatchBbs[Pstream::myProcNo()]; const treeBoundBox& tgtPatchBb = tgtPatchBbs[procI]; if (srcPatchBb.overlaps(tgtPatchBb)) { // Send over complete patch voxel map. Tbd: could // subset UOPstream os(procI, pBufs); os << srcPatchBb << patchDivisions[srcI] << patchParts[srcI]; } } } pBufs.finishedSends(); for (const int procI : Pstream::allProcs()) { if (procI != Pstream::myProcNo()) { //const treeBoundBox& srcBb = srcBbs[procI]; const treeBoundBox& srcPatchBb = srcPatchBbs[procI]; const treeBoundBox& tgtPatchBb = tgtPatchBbs[Pstream::myProcNo()]; if (srcPatchBb.overlaps(tgtPatchBb)) { UIPstream is(procI, pBufs); { treeBoundBox receivedBb(is); if (srcPatchBb != receivedBb) { FatalErrorInFunction << "proc:" << procI << " srcPatchBb:" << srcPatchBb << " receivedBb:" << receivedBb << exit(FatalError); } } const labelVector zoneDivs(is); const PackedList<2> srcPatchTypes(is); forAll(tgtCellMap, tgtCelli) { label celli = tgtCellMap[tgtCelli]; treeBoundBox cBb(cellBb(mesh_, celli)); cBb.min() -= smallVec_; cBb.max() += smallVec_; if ( overlaps ( srcPatchBb, zoneDivs, srcPatchTypes, cBb, patchCellType::PATCH ) ) { allCellTypes[celli] = HOLE; } } } } } }
The final check if a cell bounding box of meshZone A overlaps with any of the voxels defining the boundary of zellZone B is done in the following code snipplet.
bool Foam::cellCellStencils::inverseDistance::overlaps ( const boundBox& bb, const labelVector& nDivs, const PackedList<2>& vals, const treeBoundBox& subBb, const unsigned int val ) { // Checks if subBb overlaps any voxel set to val labelVector minIds(index3(bb, nDivs, subBb.min())); labelVector maxIds(index3(bb, nDivs, subBb.max())); for (direction cmpt = 0; cmpt < 3; cmpt++) { if (maxIds[cmpt] < 0 || minIds[cmpt] > nDivs[cmpt]) { return false; } } labelVector maxIndex(labelVector(nDivs[0]-1, nDivs[1]-1, nDivs[2]-1)); minIds = max(labelVector::zero, minIds); maxIds = min(maxIndex, maxIds); for (label i = minIds[0]; i <= maxIds[0]; i++) { for (label j = minIds[1]; j <= maxIds[1]; j++) { for (label k = minIds[2]; k <= maxIds[2]; k++) { label i1 = index(nDivs, labelVector(i, j, k)); if (vals[i1] == patchCellType::PATCH) { return true; } } } } return false; }
4.1.1.2 cell volume weight
The determination of the cells of meshZone A which are overlapping with boundary of the meshZone B is done over the object meshTomesh (see https://www.openfoam.com/documentation/guides/latest/api/classFoam_1_1meshToMesh.html#details). The meshToMesh objects calculates the cell-addressing between two overlapping meshes. It means if cell 100 overlaps with cells 300, 400 and 500 this relation is stored in the list srcToTgtCellAddr_. The weights (i.e. to which portion cell 100 overlaps with the cells 300, 400 and 500) is calculated with interpolation method chosen by the user. The weights are stored in the array srcToTgtCellWght_. In the case of the cell volume weight the interpolation method meshToMesh::interpolationMethod::imCellVolumeWeight is chosen when constructing the object.
Having explained the main object involved in the determination of the cells in meshZone A which are cut by the boundaries of meshZone B, we can proceed to explain the algorithm employed. First all cells in meshZone A which are adjacent to a boundary are marked as PATCH, OVERSET and OTHER. The OTHER switch is used for coupled patches like processor or cyclic. The OVERSET is used for overset patches and the PATCH switch is used used for all other patch types. This is done in the function markPatchCells
void Foam::cellCellStencils::cellVolumeWeight::markPatchCells ( const fvMesh& mesh, const labelList& cellMap, labelList& patchCellTypes ) const { const fvBoundaryMesh& pbm = mesh.boundary(); forAll(pbm, patchI) { const fvPatch& fvp = pbm[patchI]; const labelList& fc = fvp.faceCells(); if (isA<oversetFvPatch>(fvp)) { //Pout<< "Marking cells on overset patch " << fvp.name() << endl; forAll(fc, i) { label cellI = fc[i]; patchCellTypes[cellMap[cellI]] = OVERSET; } } else if (!fvPatch::constraintType(fvp.type())) { //Pout<< "Marking cells on proper patch " << fvp.name() // << " with type " << fvp.type() << endl; forAll(fc, i) { label cellI = fc[i]; if (patchCellTypes[cellMap[cellI]] != OVERSET) { patchCellTypes[cellMap[cellI]] = PATCH; } } } } }
The generation of the meshToMesh objects and the determination of the cells in meshZone A cut by the boundaries of meshZone B are done in a single loop:
for (label srcI = 0; srcI < meshParts.size()-1; srcI++)
{
const fvMesh& srcMesh = meshParts[srcI].subMesh();
const labelList& srcCellMap = meshParts[srcI].cellMap();
for (label tgtI = srcI+1; tgtI < meshParts.size(); tgtI++)
{
const fvMesh& tgtMesh = meshParts[tgtI].subMesh();
const labelList& tgtCellMap = meshParts[tgtI].cellMap();
meshToMesh mapper
(
srcMesh,
tgtMesh,
meshToMesh::interpolationMethod::imCellVolumeWeight,
HashTable<word>(0), // patchMap,
wordList(0), // cuttingPatches
meshToMesh::procMapMethod::pmAABB,
false // do not normalise
);
{
// Get tgt patch types on src mesh
labelList interpolatedTgtPatchTypes(srcMesh.nCells(), -1);
interpolatePatchTypes
(
mapper.tgtMap(), // How to get remote data local
mapper.srcToTgtCellAddr(),
labelList(labelUIndList(allPatchTypes, tgtCellMap)),
interpolatedTgtPatchTypes
);
// Get target cell labels in global cell indexing (on overall
// mesh)
labelList tgtGlobalCells(tgtMesh.nCells());
{
forAll(tgtCellMap, tgtCellI)
{
label cellI = tgtCellMap[tgtCellI];
tgtGlobalCells[tgtCellI] = globalCells.toGlobal(cellI);
}
if (mapper.tgtMap())
{
mapper.tgtMap()->distribute(tgtGlobalCells);
}
}
combineCellTypes
(
srcI,
srcMesh,
srcCellMap,
tgtI,
mapper.srcToTgtCellAddr(),
mapper.srcToTgtCellWght(),
tgtGlobalCells,
interpolatedTgtPatchTypes,
// Overall mesh data
allStencil,
allWeights,
allCellTypes,
allDonorID
);
}
{
// Get src patch types on tgt mesh
labelList interpolatedSrcPatchTypes(tgtMesh.nCells(), -1);
interpolatePatchTypes
(
mapper.srcMap(), // How to get remote data local
mapper.tgtToSrcCellAddr(),
labelList(labelUIndList(allPatchTypes, srcCellMap)),
interpolatedSrcPatchTypes
);
labelList srcGlobalCells(srcMesh.nCells());
{
forAll(srcCellMap, srcCellI)
{
label cellI = srcCellMap[srcCellI];
srcGlobalCells[srcCellI] = globalCells.toGlobal(cellI);
}
if (mapper.srcMap())
{
mapper.srcMap()->distribute(srcGlobalCells);
}
}
combineCellTypes
(
tgtI,
tgtMesh,
tgtCellMap,
srcI,
mapper.tgtToSrcCellAddr(),
mapper.tgtToSrcCellWght(),
srcGlobalCells,
interpolatedSrcPatchTypes,
// Overall mesh data
allStencil,
allWeights,
allCellTypes,
allDonorID
);
}
}
}
Next we will see in a bit more detail what is done in the previous loop. We see from the above loop, that for each combination source and target mesh a meshToMesh object is created. That means that for each cell of the source mesh the mapping to the target mesh has to be created. This is of course very time consuming and may explain why the cellVolumeWeight interpolation is much more expensive compared to other interpolation types.
Knowing the addressing between the source and target mesh (or cells of meshZone A to cells of meshZone B) determination of the cells of meshZone A which are cut be the boundaries is very easy. Let's recall that the labelList allPatchTypes stores the patch types (PATCH for a patch, OVERSET for overset patches, OTHER for coupled patches and -1 for internal cells) at the index of the cell. So know one can go through the array allPatchTypes, look if there is a PATCH, OVERSET or OTHER label contained in it. The cell index is also known at which these labels are store. Knowing the index one can look into the source to target adressing mapper.srcToTgtCellAddr to fine the cells of meshZone A which are overlapping with a boundary of meshZone B. How it is done in detail can be seen from the code snippet below:
void Foam::cellCellStencils::cellVolumeWeight::interpolatePatchTypes ( const labelListList& addressing, const labelList& patchTypes, labelList& result ) const { forAll(result, cellI) { const labelList& slots = addressing[cellI]; forAll(slots, i) { label type = patchTypes[slots[i]]; if (type == OVERSET) { // 'overset' overrides anything result[cellI] = OVERSET; break; } else if (type == PATCH) { // 'patch' overrides -1 and 'other' result[cellI] = PATCH; } else if (result[cellI] == -1) { // 'other' overrides -1 only result[cellI] = OTHER; } } } }
4.1.2 Find cells inside the region cut by the walls
Once the cells in a meshZone A cut by walls of meshZone B are marked as holes, the cells inside this region have also be marked as HOLE. This is done in the function findHoles:
void Foam::cellCellStencils::inverseDistance::findHoles ( const globalIndex& globalCells, const fvMesh& mesh, const labelList& zoneID, const labelListList& stencil, labelList& cellTypes ) const { const fvBoundaryMesh& pbm = mesh.boundary(); const labelList& own = mesh.faceOwner(); const labelList& nei = mesh.faceNeighbour(); // The input cellTypes will be // - HOLE : cell part covered by other-mesh patch // - INTERPOLATED : cell fully covered by other-mesh patch // or next to 'overset' patch // - CALCULATED : otherwise // // so we start a walk from our patches and any cell we cannot reach // (because we walk is stopped by other-mesh patch) is a hole. DebugInfo<< FUNCTION_NAME << " : Starting hole flood filling" << endl; DebugInfo<< FUNCTION_NAME << " : Starting hole cells : " << findIndices(cellTypes, HOLE).size() << endl; boolList isBlockedFace(mesh.nFaces(), false); label nBlocked = 0; for (label faceI = 0; faceI < mesh.nInternalFaces(); faceI++) { label ownType = cellTypes[own[faceI]]; label neiType = cellTypes[nei[faceI]]; if ( (ownType == HOLE && neiType != HOLE) || (ownType != HOLE && neiType == HOLE) ) { isBlockedFace[faceI] = true; nBlocked++; } } DebugInfo<< FUNCTION_NAME << " : Marked internal hole boundaries : " << nBlocked << endl; labelList nbrCellTypes; syncTools::swapBoundaryCellList(mesh, cellTypes, nbrCellTypes); for (label faceI = mesh.nInternalFaces(); faceI < mesh.nFaces(); faceI++) { label ownType = cellTypes[own[faceI]]; label neiType = nbrCellTypes[faceI-mesh.nInternalFaces()]; if ( (ownType == HOLE && neiType != HOLE) || (ownType != HOLE && neiType == HOLE) ) { isBlockedFace[faceI] = true; nBlocked++; } } DebugInfo<< FUNCTION_NAME << " : Marked all hole boundaries : " << nBlocked << endl; // Determine regions regionSplit cellRegion(mesh, isBlockedFace); const label nRegions = cellRegion.nRegions(); //labelList cellRegion; //label nRegions = -1; //{ // const globalIndex globalFaces(mesh.nFaces()); // uncompactedRegionSplit // ( // mesh, // globalFaces, // gMax(zoneID)+1, // zoneID, // cellTypes, // isBlockedFace, // cellRegion // ); // autoPtr<globalIndex> globalRegions // ( // compactedRegionSplit // ( // mesh, // globalFaces, // cellRegion // ) // ); // nRegions = globalRegions().size(); //} DebugInfo<< FUNCTION_NAME << " : Determined regions : " << nRegions << endl; //Info<< typeName << " : detected " << nRegions // << " mesh regions after overset" << nl << endl; // Now we'll have a mesh split according to where there are cells // covered by the other-side patches. See what we can reach from our // real patches // 0 : region not yet determined // 1 : borders blockage so is not ok (but can be overridden by real // patch) // 2 : has real patch in it so is reachable labelList regionType(nRegions, Zero); // See if any regions borders blockage. Note: isBlockedFace is already // parallel synchronised. { for (label faceI = 0; faceI < mesh.nInternalFaces(); faceI++) { if (isBlockedFace[faceI]) { label ownRegion = cellRegion[own[faceI]]; if (cellTypes[own[faceI]] != HOLE) { if (regionType[ownRegion] == 0) { regionType[ownRegion] = 1; } } label neiRegion = cellRegion[nei[faceI]]; if (cellTypes[nei[faceI]] != HOLE) { if (regionType[neiRegion] == 0) { regionType[neiRegion] = 1; } } } } for ( label faceI = mesh.nInternalFaces(); faceI < mesh.nFaces(); faceI++ ) { if (isBlockedFace[faceI]) { label ownRegion = cellRegion[own[faceI]]; if (regionType[ownRegion] == 0) { regionType[ownRegion] = 1; } } } } // Override with real patches forAll(pbm, patchI) { const fvPatch& fvp = pbm[patchI]; if (isA<oversetFvPatch>(fvp)) {} else if (!fvPatch::constraintType(fvp.type())) { const labelList& fc = fvp.faceCells(); forAll(fc, i) { label regionI = cellRegion[fc[i]]; if (cellTypes[fc[i]] != HOLE && regionType[regionI] != 2) { regionType[regionI] = 2; } } } } DebugInfo<< FUNCTION_NAME << " : Done local analysis" << endl; // Now we've handled // - cells next to blocked cells // - coupled boundaries // Only thing to handle is the interpolation between regions labelListList compactStencil(stencil); List<Map<label>> compactMap; mapDistribute map(globalCells, compactStencil, compactMap); DebugInfo<< FUNCTION_NAME << " : Converted stencil into compact form" << endl; while (true) { // Synchronise region status on processors // (could instead swap status through processor patches) Pstream::listCombineGather(regionType, maxEqOp<label>()); Pstream::listCombineScatter(regionType); DebugInfo<< FUNCTION_NAME << " : Gathered region type" << endl; // Communicate region status through interpolative cells labelList cellRegionType(labelUIndList(regionType, cellRegion)); map.distribute(cellRegionType); DebugInfo<< FUNCTION_NAME << " : Interpolated region type" << endl; label nChanged = 0; forAll(pbm, patchI) { const fvPatch& fvp = pbm[patchI]; if (isA<oversetFvPatch>(fvp)) { const labelUList& fc = fvp.faceCells(); forAll(fc, i) { label cellI = fc[i]; label regionI = cellRegion[cellI]; if (regionType[regionI] != 2) { const labelList& slots = compactStencil[cellI]; forAll(slots, i) { label otherType = cellRegionType[slots[i]]; if (otherType == 2) { //Pout<< "Reachable through interpolation : " // << regionI << " at cell " // << mesh.cellCentres()[cellI] << endl; regionType[regionI] = 2; nChanged++; break; } } } } } } reduce(nChanged, sumOp<label>()); DebugInfo<< FUNCTION_NAME << " : Determined regions changed : " << nChanged << endl; if (nChanged == 0) { break; } } // See which regions have not been visited (regionType == 1) forAll(cellRegion, cellI) { label type = regionType[cellRegion[cellI]]; if (type == 1 && cellTypes[cellI] != HOLE) { cellTypes[cellI] = HOLE; } } DebugInfo<< FUNCTION_NAME << " : Finished hole flood filling" << endl; }
Let's analyze a bit more in detail how the algorithm of the filling of the cells of mesh Zone A inside the region surrounded by the walls of mesh Zone B is done. The first step is to mark all faces which are in between a hole cell and a cell which is not a hole. This is achieved by using the boolList isBlockedFace. This boolList is used as input to generate the object regionSplit cellRegion(mesh, isBlockedFace);. This class separates the mesh into distinct unconnected regions, each of which is then given a label (for details see also https://www.openfoam.com/documentation/guides/latest/api/regionSplit_8H_source.html). This means that if we have two objects with walls overlying the mesh Zone A, the cells of mesh Zone A will be given one of three possible labels. The code snippet below shows how this is done.
boolList isBlockedFace(mesh.nFaces(), false);
label nBlocked = 0;
for (label faceI = 0; faceI < mesh.nInternalFaces(); faceI++)
{
label ownType = cellTypes[own[faceI]];
label neiType = cellTypes[nei[faceI]];
if
(
(ownType == HOLE && neiType != HOLE)
|| (ownType != HOLE && neiType == HOLE)
)
{
isBlockedFace[faceI] = true;
nBlocked++;
}
}
DebugInfo<< FUNCTION_NAME << " : Marked internal hole boundaries : "
<< nBlocked << endl;
labelList nbrCellTypes;
syncTools::swapBoundaryCellList(mesh, cellTypes, nbrCellTypes);
for (label faceI = mesh.nInternalFaces(); faceI < mesh.nFaces(); faceI++)
{
label ownType = cellTypes[own[faceI]];
label neiType = nbrCellTypes[faceI-mesh.nInternalFaces()];
if
(
(ownType == HOLE && neiType != HOLE)
|| (ownType != HOLE && neiType == HOLE)
)
{
isBlockedFace[faceI] = true;
nBlocked++;
}
}
DebugInfo<< FUNCTION_NAME << " : Marked all hole boundaries : "
<< nBlocked << endl;
// Determine regions
regionSplit cellRegion(mesh, isBlockedFace);
const label nRegions = cellRegion.nRegions();
Having split the mesh into different regions which are inside and outside of regions surrounded by wall, the next step is to determine which regions are inside and which are outside since the knowledge of the cell region does not imply if it is behind an object. The type of region is marked in the labelList regionType:
// Now we'll have a mesh split according to where there are cells
// covered by the other-side patches. See what we can reach from our
// real patches
// 0 : region not yet determined
// 1 : borders blockage so is not ok (but can be overridden by real
// patch)
// 2 : has real patch in it so is reachable
labelList regionType(nRegions, Zero);
4.2 Interpolated cells
Interpolated cells are those adjacent of the overset patches. Furthermore also a buffer layer of interpolated cells are places around HOLE cells. Unfortunately there are smaller differences in the function walkFront which sets this interpolation layer between the inverse distance and the cell volume weight method. The differences are however small and only the walkFront function in the cell volume weight is explained.
void Foam::cellCellStencils::cellVolumeWeight::walkFront ( const scalar layerRelax, labelList& allCellTypes, scalarField& allWeight ) const { // Current front bitSet isFront(mesh_.nFaces()); // unused: bitSet doneCell(mesh_.nCells()); const fvBoundaryMesh& fvm = mesh_.boundary(); // 'overset' patches forAll(fvm, patchI) { if (isA<oversetFvPatch>(fvm[patchI])) { //Pout<< "Storing faces on patch " << fvm[patchI].name() << endl; forAll(fvm[patchI], i) { isFront.set(fvm[patchI].start()+i); } } } // Outside of 'hole' region { const labelList& own = mesh_.faceOwner(); const labelList& nei = mesh_.faceNeighbour(); for (label faceI = 0; faceI < mesh_.nInternalFaces(); faceI++) { label ownType = allCellTypes[own[faceI]]; label neiType = allCellTypes[nei[faceI]]; if ( (ownType == HOLE && neiType != HOLE) || (ownType != HOLE && neiType == HOLE) ) { //Pout<< "Front at face:" << faceI // << " at:" << mesh_.faceCentres()[faceI] << endl; isFront.set(faceI); } } labelList nbrCellTypes; syncTools::swapBoundaryCellList(mesh_, allCellTypes, nbrCellTypes); for ( label faceI = mesh_.nInternalFaces(); faceI < mesh_.nFaces(); faceI++ ) { label ownType = allCellTypes[own[faceI]]; label neiType = nbrCellTypes[faceI-mesh_.nInternalFaces()]; if ( (ownType == HOLE && neiType != HOLE) || (ownType != HOLE && neiType == HOLE) ) { //Pout<< "Front at coupled face:" << faceI // << " at:" << mesh_.faceCentres()[faceI] << endl; isFront.set(faceI); } } } // Current interpolation fraction scalar fraction = 1.0; while (fraction > SMALL && returnReduce(isFront.count(), sumOp<label>())) { // Interpolate cells on front Info<< "Front : fraction:" << fraction << " size:" << returnReduce(isFront.count(), sumOp<label>()) << endl; bitSet newIsFront(mesh_.nFaces()); forAll(isFront, faceI) { if (isFront.test(faceI)) { label own = mesh_.faceOwner()[faceI]; if (allCellTypes[own] != HOLE) { if (allWeight[own] < fraction) { allWeight[own] = fraction; //if (debug) //{ // Pout<< " setting cell " // << mesh_.cellCentres()[own] // << " to " << fraction << endl; //} allCellTypes[own] = INTERPOLATED; newIsFront.set(mesh_.cells()[own]); } } if (mesh_.isInternalFace(faceI)) { label nei = mesh_.faceNeighbour()[faceI]; if (allCellTypes[nei] != HOLE) { if (allWeight[nei] < fraction) { allWeight[nei] = fraction; //if (debug) //{ // Pout<< " setting cell " // << mesh_.cellCentres()[nei] // << " to " << fraction << endl; //} allCellTypes[nei] = INTERPOLATED; newIsFront.set(mesh_.cells()[nei]); } } } } } syncTools::syncFaceList(mesh_, newIsFront, orEqOp<unsigned int>()); isFront.transfer(newIsFront); fraction -= layerRelax; } }
Let's have a look at the above code. First all faces which delimit a HOLE cell and a non hole cell are stored in the bitset isFront. After that the entry of the cell type at the boundary are swapped (see https://www.openfoam.com/documentation/guides/latest/api/classFoam_1_1syncTools.html#ac509eef6db47b0b87366229f2fb017f9). The swapping is required to determine also faces at the border of a hole and non-hole cell adjacent to a processor boundary. After the swapping the check if a face borders a Hole and a non-hole cell is repeated only for the boundary faces. The purpose is to check if a cell adjacent to a processor boundary boarders a hole and a non-hole cell. Finally the cells which are not a hole boarding the faces stored in isFront are set to INTERPOLATED and the weight is set to 1.
4.3 Calculated cells
The determination of the CALCULATED cells is quit simple. All cells which are not HOLE or INTERPOLATED are set as CALCULATED.
5 References
- ↑ M. Breuer, G. De Nayer, M. Münsch, T. Gallinger, and R. Wüchner, “Fluid–structure interac tion using a partitioned semi-implicit predictor–corrector coupling scheme for the application of large-eddy simulation,” Journal of Fluids and Structures, vol. 29, pp. 107–130, 2012.
- ↑ W. J. Horne and K. Mahesh, “A massively-parallel, unstructured overset method to simulate moving bodies in turbulent flows,” Journal of Computational Physics, vol. 397, p. 108790, 2019.
- ↑ 2019. J. H. Ferziger, M. Perić, and R. L. Street, Computational methods for fluid dynamics. Springer, 2002, vol. 3.
- ↑ ] M. Buchmayr, Development of Fully Implicit Block Coupled Solvers for Incompressible Transient Flows. TU-Graz, 2014.
- ↑ mechanics, vol. 33, no. 1, pp. 445–490, 2001. H. Jasak, “Dynamic mesh handling in openfoam,” in 47th AIAA aerospace sciences meeting including the new horizons forum and aerospace exposition, 2009, p. 341.
- ↑ M. Breuer, G. De Nayer, M. Münsch, T. Gallinger, and R. Wüchner, “Fluid–structure interac tion using a partitioned semi-implicit predictor–corrector coupling scheme for the application of large-eddy simulation,” Journal of Fluids and Structures, vol. 29, pp. 107–130, 2012.
- ↑ mechanics, vol. 33, no. 1, pp. 445–490, 2001. H. Jasak, “Dynamic mesh handling in openfoam,” in 47th AIAA aerospace sciences meeting including the new horizons forum and aerospace exposition, 2009, p. 341.
- ↑ M. Breuer, G. De Nayer, M. Münsch, T. Gallinger, and R. Wüchner, “Fluid–structure interac tion using a partitioned semi-implicit predictor–corrector coupling scheme for the application of large-eddy simulation,” Journal of Fluids and Structures, vol. 29, pp. 107–130, 2012.